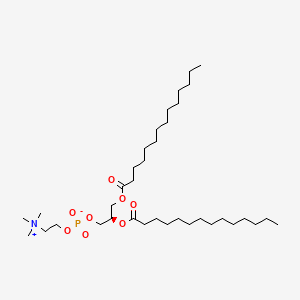
18194-24-6
18194-24-6 is a lipid of Glycerophospholipids (GP) class. 18194-24-6 is associated with abnormalities such as Cerebrovascular accident, Renal tubular disorder, Atherosclerosis, Hyperlipoproteinemia Type III and Lipid Metabolism Disorders. The involved functions are known as Process, protein folding, Catalyst, Biochemical Pathway and Fold in Medical Device Material. 18194-24-6 often locates in Tissue membrane, Membrane, periplasm, vesicle membrane and outer membrane. The associated genes with 18194-24-6 are Integral Membrane Proteins, Protein Structure, RTN4 gene, RTN4R gene and Protein, Organized by Structure. The related lipids are Micelles, dimyristoylphosphatidylglycerol, 1,2-dihexadecyl-sn-glycero-3-phosphocholine, Unilamellar Vesicles and cholesteryl oleate. The related experimental models are Mouse Model, Arthritis, Adjuvant-Induced, Disease model and Xenograft Model.
Cross Reference
Introduction
To understand associated biological information of 18194-24-6, we collected biological information of abnormalities, associated pathways, cellular/molecular locations, biological functions, related genes/proteins, lipids and common seen animal/experimental models with organized paragraphs from literatures.
What diseases are associated with 18194-24-6?
18194-24-6 is suspected in Atherosclerosis, Cardiovascular Diseases, Dehydration, Abnormal shape, Renal tubular disorder, Hyperlipoproteinemia Type III and other diseases in descending order of the highest number of associated sentences.
Related references are mostly published in these journals:
| Disease | Cross reference | Weighted score | Related literature |
|---|
Possible diseases from mapped MeSH terms on references
We collected disease MeSH terms mapped to the references associated with 18194-24-6
PubChem Associated disorders and diseases
What pathways are associated with 18194-24-6
There are no associated biomedical information in the current reference collection.
PubChem Biomolecular Interactions and Pathways
Link to PubChem Biomolecular Interactions and PathwaysWhat cellular locations are associated with 18194-24-6?
Visualization in cellular structure
Associated locations are in red color. Not associated locations are in black.
Related references are published most in these journals:
| Location | Cross reference | Weighted score | Related literatures |
|---|
What functions are associated with 18194-24-6?
Related references are published most in these journals:
| Function | Cross reference | Weighted score | Related literatures |
|---|
What lipids are associated with 18194-24-6?
Related references are published most in these journals:
| Lipid concept | Cross reference | Weighted score | Related literatures |
|---|
What genes are associated with 18194-24-6?
Related references are published most in these journals:
| Gene | Cross reference | Weighted score | Related literatures |
|---|
What common seen animal models are associated with 18194-24-6?
Mouse Model
Mouse Model are used in the study 'Association of a model class A (apolipoprotein) amphipathic alpha helical peptide with lipid: high resolution NMR studies of peptide.lipid discoidal complexes.' (Mishra VK et al., 2006).
Arthritis, Adjuvant-Induced
Arthritis, Adjuvant-Induced are used in the study 'T cell antigen receptor peptide-lipid membrane interactions using surface plasmon resonance.' (Bender V et al., 2004).
Disease model
Disease model are used in the study 'Kupffer cells do not play a role in governing the efficacy of liposomal mitoxantrone used to treat a tumor model designed to assess drug delivery to liver.' (Lim HJ et al., 2000).
Related references are published most in these journals:
| Model | Cross reference | Weighted score | Related literatures |
|---|
NCBI Entrez Crosslinks
All references with 18194-24-6
Download all related citations| Authors | Title | Published | Journal | PubMed Link |
|---|---|---|---|---|
| Dominguez GA et al. | Measurement of the bending elastic modulus in unilamellar vesicles membranes by fast field cycling NMR relaxometry. | 2016 | Chem. Phys. Lipids | pmid:27816433 |
| Scheidelaar S et al. | Effect of Polymer Composition and pH on Membrane Solubilization by Styrene-Maleic Acid Copolymers. | 2016 | Biophys. J. | pmid:27806279 |
| Gobrogge CA et al. | Temperature Dependent Solvation and Partitioning of Coumarin 152 in Phospholipid Membranes. | 2016 | J Phys Chem B | pmid:26624521 |
| Davis JH and Komljenović I | Nuclear Overhauser effect as a probe of molecular structure, dynamics and order of axially reorienting molecules in membranes. | 2016 | Biochim. Biophys. Acta | pmid:26607012 |
| Muhanna N et al. | Multimodal Image-Guided Surgical and Photodynamic Interventions in Head and Neck Cancer: From Primary Tumor to Metastatic Drainage. | 2016 | Clin. Cancer Res. | pmid:26463705 |
| Yoshimura H et al. | Two-Dimensional Crystallization of P22 Virus-Like Particles. | 2016 | J Phys Chem B | pmid:27125277 |
| Adhikari U et al. | Properties of Poloxamer Molecules and Poloxamer Micelles Dissolved in Water and Next to Lipid Bilayers: Results from Computer Simulations. | 2016 | J Phys Chem B | pmid:26719970 |
| Alsop RJ et al. | Swelling of phospholipid membranes by divalent metal ions depends on the location of the ions in the bilayers. | 2016 | Soft Matter | pmid:27453289 |
| Zhernenkov M et al. | Revealing the mechanism of passive transport in lipid bilayers via phonon-mediated nanometre-scale density fluctuations. | 2016 | Nat Commun | pmid:27175859 |
| Hasan IY and Mechler A | Cholesterol Rich Domains Identified in Unilamellar Supported Biomimetic Membranes via Nano-Viscosity Measurements. | 2016 | Anal. Chem. | pmid:27137411 |
| Madrid E and Horswell SL | Effect of Deuteration on Phase Behavior of Supported Phospholipid Bilayers: A Spectroelectrochemical Study. | 2015 | Langmuir | pmid:26536482 |
| Tanaka M et al. | Preparation and Characterization of Reconstituted Lipid-Synthetic Polymer Discoidal Particles. | 2015 | Langmuir | pmid:26531224 |
| Wölk C et al. | Investigation of Binary Lipid Mixtures of a Three-Chain Cationic Lipid with Phospholipids Suitable for Gene Delivery. | 2015 | Bioconjug. Chem. | pmid:26471337 |
| Björnerås J et al. | Analysing DHPC/DMPC bicelles by diffusion NMR and multivariate decomposition. | 2015 | Biochim. Biophys. Acta | pmid:26341141 |
| Malishev R et al. | Toxicity inhibitors protect lipid membranes from disruption by Aβ42. | 2015 | ACS Chem Neurosci | pmid:26317327 |
| Ding Y et al. | Influence of the lipid membrane environment on structure and activity of the outer membrane protein Ail from Yersinia pestis. | 2015 | Biochim. Biophys. Acta | pmid:25433311 |
| Smrt ST et al. | The influenza hemagglutinin fusion domain is an amphipathic helical hairpin that functions by inducing membrane curvature. | 2015 | J. Biol. Chem. | pmid:25398882 |
| Vad BS et al. | Phospholipid Ether Linkages Significantly Modulate the Membrane Affinity of the Antimicrobial Peptide Novicidin. | 2015 | J. Membr. Biol. | pmid:25801603 |
| Shen H et al. | An anisotropic coarse-grained model based on Gay-Berne and electric multipole potentials and its application to simulate a DMPC bilayer in an implicit solvent model. | 2015 | J Comput Chem | pmid:25788250 |
| Madej BD et al. | A Parameterization of Cholesterol for Mixed Lipid Bilayer Simulation within the Amber Lipid14 Force Field. | 2015 | J Phys Chem B | pmid:26359797 |