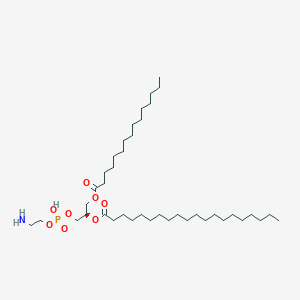
PE(15:0/20:0)
PE(15:0/20:0) is a lipid of Glycerophospholipids (GP) class. Pe(15:0/20:0) is associated with abnormalities such as Exanthema, Infection, Painful Bladder Syndrome, Obesity and Fatty Liver. The involved functions are known as conjugation, Transcription, Genetic, Sinking, Autophagy and Protein Biosynthesis. Pe(15:0/20:0) often locates in membrane fraction, soluble, Membrane, Body tissue and Tissue membrane. The associated genes with PE(15:0/20:0) are GABARAPL2 gene, ATG10 gene, ATG12 gene, SLC33A1 gene and GABARAP gene. The related lipids are Liposomes, Lipopolysaccharides, Phosphatidylserines, Membrane Lipids and Cardiolipins. The related experimental models are Knock-out and Cancer Model.
Cross Reference
Introduction
To understand associated biological information of PE(15:0/20:0), we collected biological information of abnormalities, associated pathways, cellular/molecular locations, biological functions, related genes/proteins, lipids and common seen animal/experimental models with organized paragraphs from literatures.
What diseases are associated with PE(15:0/20:0)?
PE(15:0/20:0) is suspected in Infection, CONE-ROD DYSTROPHY 1 (disorder), Diabetes, Obesity, Malaria, Atherosclerosis and other diseases in descending order of the highest number of associated sentences.
Related references are mostly published in these journals:
| Disease | Cross reference | Weighted score | Related literature |
|---|
Possible diseases from mapped MeSH terms on references
We collected disease MeSH terms mapped to the references associated with PE(15:0/20:0)
PubChem Associated disorders and diseases
What pathways are associated with PE(15:0/20:0)
There are no associated biomedical information in the current reference collection.
PubChem Biomolecular Interactions and Pathways
Link to PubChem Biomolecular Interactions and PathwaysWhat cellular locations are associated with PE(15:0/20:0)?
Visualization in cellular structure
Associated locations are in red color. Not associated locations are in black.
Related references are published most in these journals:
| Location | Cross reference | Weighted score | Related literatures |
|---|
What functions are associated with PE(15:0/20:0)?
Related references are published most in these journals:
| Function | Cross reference | Weighted score | Related literatures |
|---|
What lipids are associated with PE(15:0/20:0)?
Related references are published most in these journals:
| Lipid concept | Cross reference | Weighted score | Related literatures |
|---|
What genes are associated with PE(15:0/20:0)?
Related references are published most in these journals:
| Gene | Cross reference | Weighted score | Related literatures |
|---|
What common seen animal models are associated with PE(15:0/20:0)?
Knock-out
Knock-out are used in the study 'Sequential synthesis and methylation of phosphatidylethanolamine promote lipid droplet biosynthesis and stability in tissue culture and in vivo.' (Hörl G et al., 2011) and Knock-out are used in the study 'An Atg4B mutant hampers the lipidation of LC3 paralogues and causes defects in autophagosome closure.' (Fujita N et al., 2008).
Cancer Model
Cancer Model are used in the study 'Improving penetration in tumors with nanoassemblies of phospholipids and doxorubicin.' (Tang N et al., 2007).
Related references are published most in these journals:
| Model | Cross reference | Weighted score | Related literatures |
|---|
NCBI Entrez Crosslinks
All references with PE(15:0/20:0)
Download all related citations| Authors | Title | Published | Journal | PubMed Link |
|---|---|---|---|---|
| Park M et al. | Flavobacterium chuncheonense sp. nov. and Flavobacterium luteum sp. nov., isolated from a freshwater lake. | 2017 | Int. J. Syst. Evol. Microbiol. | pmid:28920853 |
| Cho ES et al. | Flavimarina flava sp. nov., isolated from Salicornia herbacea. | 2017 | Int. J. Syst. Evol. Microbiol. | pmid:28920849 |
| Cao M et al. | Edaphobaculum flavum gen. nov., sp. nov., a member of family Chitinophagaceae, isolated from grassland soil. | 2017 | Int. J. Syst. Evol. Microbiol. | pmid:28920838 |
| Frasson D et al. | Pseudomonas wadenswilerensis sp. nov. and Pseudomonas reidholzensis sp. nov., two novel species within the Pseudomonas putida group isolated from forest soil. | 2017 | Int. J. Syst. Evol. Microbiol. | pmid:28820109 |
| Park S et al. | Umboniibacter caenipelagi sp. nov., isolated from a tidal flat. | 2017 | Int. J. Syst. Evol. Microbiol. | pmid:28820102 |
| Lee KC et al. | Mucilaginibacter craterilacus sp. nov., isolated from sediment soil of a crater lake. | 2017 | Int. J. Syst. Evol. Microbiol. | pmid:28820098 |
| Muramatsu Y et al. | Reclassification of Flexibacter tractuosus NBRC 15981T as Marivirga harenae sp. nov. in the family Flammeovirgaceae. | 2017 | Int. J. Syst. Evol. Microbiol. | pmid:28646633 |
| SedláÄek I et al. | Red-pink pigmented Hymenobacter coccineus sp. nov., Hymenobacter lapidarius sp. nov. and Hymenobacter glacialis sp. nov., isolated from rocks in Antarctica. | 2017 | Int. J. Syst. Evol. Microbiol. | pmid:28629503 |
| Lewenza S et al. | Hyperbiofilm phenotype of Pseudomonas aeruginosa defective for the PlcB and PlcN secreted phospholipases. | 2017 | Can. J. Microbiol. | pmid:28609638 |
| Ben Mouhoub R et al. | Unraveling the expression of genes involved in the biosynthesis pathway of cardiolipin and phosphatidylethanolamine in Salmonella Hadar grown under static magnetic field 200Â mT. | 2017 | Microb. Pathog. | pmid:28923603 |
| Turner KM et al. | Changes in Lipids and Inflammatory Markers after Consuming Diets High in Red Meat or Dairy for Four Weeks. | 2017 | Nutrients | pmid:28817063 |
| Cerminati S et al. | Development of a highly efficient oil degumming process using a novel phosphatidylinositol-specific phospholipase C enzyme. | 2017 | Appl. Microbiol. Biotechnol. | pmid:28238084 |
| Elvas F et al. | Phosphatidylethanolamine targeting for cell death imaging in early treatment response evaluation and disease diagnosis. | 2017 | Apoptosis | pmid:28623512 |
| Zhang YJ et al. | Euzebyella marina sp. nov., isolated from seawater. | 2017 | Int. J. Syst. Evol. Microbiol. | pmid:27911255 |
| Kang H et al. | Polaribacter lacunae sp. nov., isolated from a lagoon. | 2017 | Int. J. Syst. Evol. Microbiol. | pmid:27902370 |
| Kim MC et al. | Hymenobacter rutilus sp. nov., isolated from marine sediment in the Arctic. | 2017 | Int. J. Syst. Evol. Microbiol. | pmid:27902346 |
| Panschin I et al. | Description of Gramella forsetii sp. nov., a marine Flavobacteriaceae isolated from North Sea water, and emended description of Gramella gaetbulicola Cho et al. 2011. | 2017 | Int. J. Syst. Evol. Microbiol. | pmid:27902319 |
| Unsay JD et al. | Pro-apoptotic cBid and Bax exhibit distinct membrane remodeling activities: An AFM study. | 2017 | Biochim. Biophys. Acta | pmid:27755971 |
| Vazdar K et al. | Reaction Mechanism of Covalent Modification of Phosphatidylethanolamine Lipids by Reactive Aldehydes 4-Hydroxy-2-nonenal and 4-Oxo-2-nonenal. | 2017 | Chem. Res. Toxicol. | pmid:28222263 |
| Dimmer KS and Rapaport D | Mitochondrial contact sites as platforms for phospholipid exchange. | 2017 | Biochim. Biophys. Acta | pmid:27477677 |