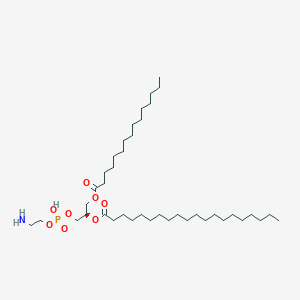
PE(15:0/20:0)
PE(15:0/20:0) is a lipid of Glycerophospholipids (GP) class. Pe(15:0/20:0) is associated with abnormalities such as Exanthema, Infection, Painful Bladder Syndrome, Obesity and Fatty Liver. The involved functions are known as conjugation, Transcription, Genetic, Sinking, Autophagy and Protein Biosynthesis. Pe(15:0/20:0) often locates in membrane fraction, soluble, Membrane, Body tissue and Tissue membrane. The associated genes with PE(15:0/20:0) are GABARAPL2 gene, ATG10 gene, ATG12 gene, SLC33A1 gene and GABARAP gene. The related lipids are Liposomes, Lipopolysaccharides, Phosphatidylserines, Membrane Lipids and Cardiolipins. The related experimental models are Knock-out and Cancer Model.
Cross Reference
Introduction
To understand associated biological information of PE(15:0/20:0), we collected biological information of abnormalities, associated pathways, cellular/molecular locations, biological functions, related genes/proteins, lipids and common seen animal/experimental models with organized paragraphs from literatures.
What diseases are associated with PE(15:0/20:0)?
PE(15:0/20:0) is suspected in Infection, CONE-ROD DYSTROPHY 1 (disorder), Diabetes, Obesity, Malaria, Atherosclerosis and other diseases in descending order of the highest number of associated sentences.
Related references are mostly published in these journals:
| Disease | Cross reference | Weighted score | Related literature |
|---|
Possible diseases from mapped MeSH terms on references
We collected disease MeSH terms mapped to the references associated with PE(15:0/20:0)
PubChem Associated disorders and diseases
What pathways are associated with PE(15:0/20:0)
There are no associated biomedical information in the current reference collection.
PubChem Biomolecular Interactions and Pathways
Link to PubChem Biomolecular Interactions and PathwaysWhat cellular locations are associated with PE(15:0/20:0)?
Visualization in cellular structure
Associated locations are in red color. Not associated locations are in black.
Related references are published most in these journals:
| Location | Cross reference | Weighted score | Related literatures |
|---|
What functions are associated with PE(15:0/20:0)?
Related references are published most in these journals:
| Function | Cross reference | Weighted score | Related literatures |
|---|
What lipids are associated with PE(15:0/20:0)?
Related references are published most in these journals:
| Lipid concept | Cross reference | Weighted score | Related literatures |
|---|
What genes are associated with PE(15:0/20:0)?
Related references are published most in these journals:
| Gene | Cross reference | Weighted score | Related literatures |
|---|
What common seen animal models are associated with PE(15:0/20:0)?
Knock-out
Knock-out are used in the study 'Sequential synthesis and methylation of phosphatidylethanolamine promote lipid droplet biosynthesis and stability in tissue culture and in vivo.' (Hörl G et al., 2011) and Knock-out are used in the study 'An Atg4B mutant hampers the lipidation of LC3 paralogues and causes defects in autophagosome closure.' (Fujita N et al., 2008).
Cancer Model
Cancer Model are used in the study 'Improving penetration in tumors with nanoassemblies of phospholipids and doxorubicin.' (Tang N et al., 2007).
Related references are published most in these journals:
| Model | Cross reference | Weighted score | Related literatures |
|---|
NCBI Entrez Crosslinks
All references with PE(15:0/20:0)
Download all related citations| Authors | Title | Published | Journal | PubMed Link |
|---|---|---|---|---|
| Nair U et al. | A role for Atg8-PE deconjugation in autophagosome biogenesis. | 2012 | Autophagy | pmid:22622160 |
| Alirezaei M et al. | Coxsackievirus can exploit LC3 in both autophagy-dependent and -independent manners in vivo. | 2015 | Autophagy | pmid:26090585 |
| Kuma A et al. | LC3, an autophagosome marker, can be incorporated into protein aggregates independent of autophagy: caution in the interpretation of LC3 localization. | 2007 Jul-Aug | Autophagy | pmid:17387262 |
| Mizushima N and Yoshimori T | How to interpret LC3 immunoblotting. | 2007 Nov-Dec | Autophagy | pmid:17611390 |
| Yu ZQ et al. | Dual roles of Atg8-PE deconjugation by Atg4 in autophagy. | 2012 | Autophagy | pmid:22652539 |
| Mitroi DN et al. | SGPL1 (sphingosine phosphate lyase 1) modulates neuronal autophagy via phosphatidylethanolamine production. | 2017 | Autophagy | pmid:28521611 |
| Nivon M et al. | Autophagy activation by NFkappaB is essential for cell survival after heat shock. | 2009 | Autophagy | pmid:19502777 |
| Shao Y et al. | Stimulation of ATG12-ATG5 conjugation by ribonucleic acid. | 2007 Jan-Feb | Autophagy | pmid:16963840 |
| Nakatogawa H et al. | Lipidation of Atg8: how is substrate specificity determined without a canonical E3 enzyme? | 2008 | Autophagy | pmid:18690009 |
| Park JM et al. | The ULK1 complex mediates MTORC1 signaling to the autophagy initiation machinery via binding and phosphorylating ATG14. | 2016 | Autophagy | pmid:27046250 |
| Yu CY et al. | Membrane glycerolipid equilibrium under endoplasmic reticulum stress in Arabidopsis thaliana. | 2018 | Biochem. Biophys. Res. Commun. | pmid:29524407 |
| Takagi K et al. | Involvement of Golgi-associated retrograde protein complex in the recycling of the putative Dnf aminophospholipid flippases in yeast. | 2012 | Biochem. Biophys. Res. Commun. | pmid:22177957 |
| Zhang Y et al. | Crystal structure of the receptor binding domain of the botulinum C-D mosaic neurotoxin reveals potential roles of lysines 1118 and 1136 in membrane interactions. | 2011 | Biochem. Biophys. Res. Commun. | pmid:21130733 |
| Arvind TA and Rangarajan PN | Mouse Apolipoprotein L9 is a phosphatidylethanolamine-binding protein. | 2016 | Biochem. Biophys. Res. Commun. | pmid:27697524 |
| Kim YA et al. | Lipophilicity of flavonoid complexes with iron(II) and their interaction with liposomes. | 2013 | Biochem. Biophys. Res. Commun. | pmid:23357424 |
| Kegel KB et al. | Polyglutamine expansion in huntingtin increases its insertion into lipid bilayers. | 2009 | Biochem. Biophys. Res. Commun. | pmid:19607813 |
| Separovic D et al. | Ceramide response post-photodamage is absent after treatment with HA14-1. | 2006 | Biochem. Biophys. Res. Commun. | pmid:16701558 |
| Tian S et al. | Human CTP:phosphoethanolamine cytidylyltransferase: enzymatic properties and unequal catalytic roles of CTP-binding motifs in two cytidylyltransferase domains. | 2014 | Biochem. Biophys. Res. Commun. | pmid:24802409 |
| Kaliszewski P et al. | Enhanced levels of Pis1p (phosphatidylinositol synthase) improve the growth of Saccharomyces cerevisiae cells deficient in Rsp5 ubiquitin ligase. | 2006 | Biochem. J. | pmid:16363994 |
| Maheshwari S et al. | Biochemical characterization of Plasmodium falciparum CTP:phosphoethanolamine cytidylyltransferase shows that only one of the two cytidylyltransferase domains is active. | 2013 | Biochem. J. | pmid:23198904 |