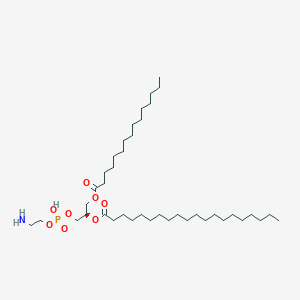
PE(15:0/20:0)
PE(15:0/20:0) is a lipid of Glycerophospholipids (GP) class. Pe(15:0/20:0) is associated with abnormalities such as Exanthema, Infection, Painful Bladder Syndrome, Obesity and Fatty Liver. The involved functions are known as conjugation, Transcription, Genetic, Sinking, Autophagy and Protein Biosynthesis. Pe(15:0/20:0) often locates in membrane fraction, soluble, Membrane, Body tissue and Tissue membrane. The associated genes with PE(15:0/20:0) are GABARAPL2 gene, ATG10 gene, ATG12 gene, SLC33A1 gene and GABARAP gene. The related lipids are Liposomes, Lipopolysaccharides, Phosphatidylserines, Membrane Lipids and Cardiolipins. The related experimental models are Knock-out and Cancer Model.
Cross Reference
Introduction
To understand associated biological information of PE(15:0/20:0), we collected biological information of abnormalities, associated pathways, cellular/molecular locations, biological functions, related genes/proteins, lipids and common seen animal/experimental models with organized paragraphs from literatures.
What diseases are associated with PE(15:0/20:0)?
PE(15:0/20:0) is suspected in Infection, CONE-ROD DYSTROPHY 1 (disorder), Diabetes, Obesity, Malaria, Atherosclerosis and other diseases in descending order of the highest number of associated sentences.
Related references are mostly published in these journals:
| Disease | Cross reference | Weighted score | Related literature |
|---|
Possible diseases from mapped MeSH terms on references
We collected disease MeSH terms mapped to the references associated with PE(15:0/20:0)
PubChem Associated disorders and diseases
What pathways are associated with PE(15:0/20:0)
There are no associated biomedical information in the current reference collection.
PubChem Biomolecular Interactions and Pathways
Link to PubChem Biomolecular Interactions and PathwaysWhat cellular locations are associated with PE(15:0/20:0)?
Visualization in cellular structure
Associated locations are in red color. Not associated locations are in black.
Related references are published most in these journals:
| Location | Cross reference | Weighted score | Related literatures |
|---|
What functions are associated with PE(15:0/20:0)?
Related references are published most in these journals:
| Function | Cross reference | Weighted score | Related literatures |
|---|
What lipids are associated with PE(15:0/20:0)?
Related references are published most in these journals:
| Lipid concept | Cross reference | Weighted score | Related literatures |
|---|
What genes are associated with PE(15:0/20:0)?
Related references are published most in these journals:
| Gene | Cross reference | Weighted score | Related literatures |
|---|
What common seen animal models are associated with PE(15:0/20:0)?
Knock-out
Knock-out are used in the study 'Sequential synthesis and methylation of phosphatidylethanolamine promote lipid droplet biosynthesis and stability in tissue culture and in vivo.' (Hörl G et al., 2011) and Knock-out are used in the study 'An Atg4B mutant hampers the lipidation of LC3 paralogues and causes defects in autophagosome closure.' (Fujita N et al., 2008).
Cancer Model
Cancer Model are used in the study 'Improving penetration in tumors with nanoassemblies of phospholipids and doxorubicin.' (Tang N et al., 2007).
Related references are published most in these journals:
| Model | Cross reference | Weighted score | Related literatures |
|---|
NCBI Entrez Crosslinks
All references with PE(15:0/20:0)
Download all related citations| Authors | Title | Published | Journal | PubMed Link |
|---|---|---|---|---|
| Sullivan EM et al. | Docosahexaenoic acid lowers cardiac mitochondrial enzyme activity by replacing linoleic acid in the phospholipidome. | 2018 | J. Biol. Chem. | pmid:29162722 |
| Cui MD et al. | Pedobacter agrisoli sp. nov., isolated from farmland soil. | 2018 | Int. J. Syst. Evol. Microbiol. | pmid:29458546 |
| Jiang WK et al. | Terrimonas soli sp. nov., isolated from farmland soil. | 2018 | Int. J. Syst. Evol. Microbiol. | pmid:29458527 |
| Choi JY et al. | Flavobacterium kingsejongi sp. nov., a carotenoid-producing species isolated from Antarctic penguin faeces. | 2018 | Int. J. Syst. Evol. Microbiol. | pmid:29458488 |
| Nedashkovskaya OI et al. | Aquimarina algiphila sp. nov., a chitin degrading bacterium isolated from the red alga Tichocarpus crinitus. | 2018 | Int. J. Syst. Evol. Microbiol. | pmid:29458485 |
| Zhang B et al. | Pedobacter quisquiliarum sp. nov., isolated from activated sludge. | 2018 | Int. J. Syst. Evol. Microbiol. | pmid:29231155 |
| Osman G et al. | Pontibacter brevis sp. nov., isolated from rhizosphere soil of Tamarix ramosissima. | 2018 | Int. J. Syst. Evol. Microbiol. | pmid:29205133 |
| Park S et al. | Tenacibaculum insulae sp. nov., isolated from a tidal flat. | 2018 | Int. J. Syst. Evol. Microbiol. | pmid:29148365 |
| Pleguezuelos-Villa M et al. | A novel ultradeformable liposomes of Naringin for anti-inflammatory therapy. | 2018 | Colloids Surf B Biointerfaces | pmid:29216513 |
| Yu WN et al. | Agaribacter flavus sp. nov., isolated from red algae. | 2018 | Int. J. Syst. Evol. Microbiol. | pmid:30091699 |
| Lee DW et al. | Oceanimonas marisflavi sp. nov., a polycyclic aromatic hydrocarbon-degrading marine bacterium. | 2018 | Int. J. Syst. Evol. Microbiol. | pmid:30040062 |
| Ohn JE et al. | Hymenobacter rufus sp. nov., a bacterium isolated from soil. | 2018 | Int. J. Syst. Evol. Microbiol. | pmid:30028287 |
| Chhetri G et al. | Edaphorhabdus rosea gen. nov., sp. nov., a new member of the family Cytophagaceae isolated from soil in South Korea. | 2018 | Antonie Van Leeuwenhoek | pmid:30027519 |
| Chen H et al. | Hymenobacter bucti sp. nov., isolated from subsurface sandstone sediment. | 2018 | Int. J. Syst. Evol. Microbiol. | pmid:30024374 |
| Kim SY et al. | Flavobacterium knui sp. nov., a novel member of the family Flavobacteriaceae. | 2018 | Antonie Van Leeuwenhoek | pmid:30022265 |
| Kristyanto S et al. | Characterization of Flavobacterium aquimarinum sp. nov., a halotolerant bacterium isolated from seawater. | 2018 | J. Microbiol. | pmid:29721828 |
| Chen Q et al. | Flavobacterium zaozhuangense sp. nov., a new member of the family Flavobacteriaceae, isolated from metolachlor-contaminated soil. | 2018 | Antonie Van Leeuwenhoek | pmid:29713912 |
| May-Zhang LS et al. | Modification by isolevuglandins, highly reactive γ-ketoaldehydes, deleteriously alters high-density lipoprotein structure and function. | 2018 | J. Biol. Chem. | pmid:29712723 |
| Lee Y and Jeon CO | Siphonobacter curvatus sp. nov., isolated from a freshwater river. | 2018 | Int. J. Syst. Evol. Microbiol. | pmid:29676722 |
| Lee Y and Jeon CO | Flavobacterium alvei sp. nov., isolated from a freshwater river. | 2018 | Int. J. Syst. Evol. Microbiol. | pmid:29676721 |