| MeSH term | MeSH ID | Detail |
|---|---|---|
| Venous Thromboembolism | D054556 | 2 associated lipids |
| Barth Syndrome | D056889 | 3 associated lipids |
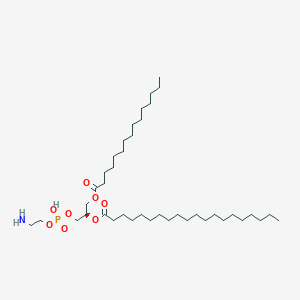
PE(15:0/20:0)
PE(15:0/20:0) is a lipid of Glycerophospholipids (GP) class. Pe(15:0/20:0) is associated with abnormalities such as Exanthema, Infection, Painful Bladder Syndrome, Obesity and Fatty Liver. The involved functions are known as conjugation, Transcription, Genetic, Sinking, Autophagy and Protein Biosynthesis. Pe(15:0/20:0) often locates in membrane fraction, soluble, Membrane, Body tissue and Tissue membrane. The associated genes with PE(15:0/20:0) are GABARAPL2 gene, ATG10 gene, ATG12 gene, SLC33A1 gene and GABARAP gene. The related lipids are Liposomes, Lipopolysaccharides, Phosphatidylserines, Membrane Lipids and Cardiolipins. The related experimental models are Knock-out and Cancer Model.
Cross Reference
Introduction
To understand associated biological information of PE(15:0/20:0), we collected biological information of abnormalities, associated pathways, cellular/molecular locations, biological functions, related genes/proteins, lipids and common seen animal/experimental models with organized paragraphs from literatures.
What diseases are associated with PE(15:0/20:0)?
PE(15:0/20:0) is suspected in Infection, CONE-ROD DYSTROPHY 1 (disorder), Diabetes, Obesity, Malaria, Atherosclerosis and other diseases in descending order of the highest number of associated sentences.
Related references are mostly published in these journals:
| Disease | Cross reference | Weighted score | Related literature |
|---|
Possible diseases from mapped MeSH terms on references
We collected disease MeSH terms mapped to the references associated with PE(15:0/20:0)
PubChem Associated disorders and diseases
What pathways are associated with PE(15:0/20:0)
There are no associated biomedical information in the current reference collection.
PubChem Biomolecular Interactions and Pathways
Link to PubChem Biomolecular Interactions and PathwaysWhat cellular locations are associated with PE(15:0/20:0)?
Visualization in cellular structure
Associated locations are in red color. Not associated locations are in black.
Related references are published most in these journals:
| Location | Cross reference | Weighted score | Related literatures |
|---|
What functions are associated with PE(15:0/20:0)?
Related references are published most in these journals:
| Function | Cross reference | Weighted score | Related literatures |
|---|
What lipids are associated with PE(15:0/20:0)?
Related references are published most in these journals:
| Lipid concept | Cross reference | Weighted score | Related literatures |
|---|
What genes are associated with PE(15:0/20:0)?
Related references are published most in these journals:
| Gene | Cross reference | Weighted score | Related literatures |
|---|
What common seen animal models are associated with PE(15:0/20:0)?
Knock-out
Knock-out are used in the study 'Sequential synthesis and methylation of phosphatidylethanolamine promote lipid droplet biosynthesis and stability in tissue culture and in vivo.' (Hörl G et al., 2011) and Knock-out are used in the study 'An Atg4B mutant hampers the lipidation of LC3 paralogues and causes defects in autophagosome closure.' (Fujita N et al., 2008).
Cancer Model
Cancer Model are used in the study 'Improving penetration in tumors with nanoassemblies of phospholipids and doxorubicin.' (Tang N et al., 2007).
Related references are published most in these journals:
| Model | Cross reference | Weighted score | Related literatures |
|---|
NCBI Entrez Crosslinks
All references with PE(15:0/20:0)
Download all related citations| Authors | Title | Published | Journal | PubMed Link |
|---|---|---|---|---|
| Li W et al. | Spirosoma horti sp. nov., isolated from apple orchard soil. | 2018 | Int. J. Syst. Evol. Microbiol. | pmid:29458662 |
| Cui MD et al. | Pedobacter agrisoli sp. nov., isolated from farmland soil. | 2018 | Int. J. Syst. Evol. Microbiol. | pmid:29458546 |
| Jiang WK et al. | Terrimonas soli sp. nov., isolated from farmland soil. | 2018 | Int. J. Syst. Evol. Microbiol. | pmid:29458527 |
| Nahar S and Cha CJ | Paenibacillus limicola sp. nov., isolated from tidal flat sediment. | 2018 | Int. J. Syst. Evol. Microbiol. | pmid:29235981 |
| Zhang B et al. | Pedobacter quisquiliarum sp. nov., isolated from activated sludge. | 2018 | Int. J. Syst. Evol. Microbiol. | pmid:29231155 |
| Králová S et al. | Flavobacterium chryseum sp. nov. and Flavobacterium psychroterrae sp. nov., novel environmental bacteria isolated from Antarctica. | 2018 | Int. J. Syst. Evol. Microbiol. | pmid:30095387 |
| Yu WN et al. | Agaribacter flavus sp. nov., isolated from red algae. | 2018 | Int. J. Syst. Evol. Microbiol. | pmid:30091699 |
| Lee DW et al. | Oceanimonas marisflavi sp. nov., a polycyclic aromatic hydrocarbon-degrading marine bacterium. | 2018 | Int. J. Syst. Evol. Microbiol. | pmid:30040062 |
| Dahal RH and Kim J | Dyadobacter flavus sp. nov. and Dyadobacter terricola sp. nov., two novel members of the family Cytophagaceae isolated from forest soil. | 2018 | Arch. Microbiol. | pmid:29737366 |
| Jin MF et al. | Terrabacter ginsengisoli sp. nov., isolated from ginseng cultivating soil. | 2018 | J. Microbiol. | pmid:29721830 |