| MeSH term | MeSH ID | Detail |
|---|---|---|
| Venous Thromboembolism | D054556 | 2 associated lipids |
| Barth Syndrome | D056889 | 3 associated lipids |
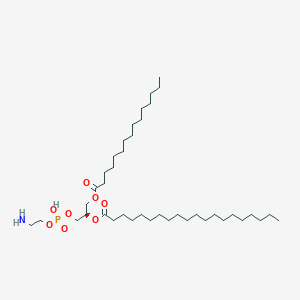
PE(15:0/20:0)
PE(15:0/20:0) is a lipid of Glycerophospholipids (GP) class. Pe(15:0/20:0) is associated with abnormalities such as Exanthema, Infection, Painful Bladder Syndrome, Obesity and Fatty Liver. The involved functions are known as conjugation, Transcription, Genetic, Sinking, Autophagy and Protein Biosynthesis. Pe(15:0/20:0) often locates in membrane fraction, soluble, Membrane, Body tissue and Tissue membrane. The associated genes with PE(15:0/20:0) are GABARAPL2 gene, ATG10 gene, ATG12 gene, SLC33A1 gene and GABARAP gene. The related lipids are Liposomes, Lipopolysaccharides, Phosphatidylserines, Membrane Lipids and Cardiolipins. The related experimental models are Knock-out and Cancer Model.
Cross Reference
Introduction
To understand associated biological information of PE(15:0/20:0), we collected biological information of abnormalities, associated pathways, cellular/molecular locations, biological functions, related genes/proteins, lipids and common seen animal/experimental models with organized paragraphs from literatures.
What diseases are associated with PE(15:0/20:0)?
PE(15:0/20:0) is suspected in Infection, CONE-ROD DYSTROPHY 1 (disorder), Diabetes, Obesity, Malaria, Atherosclerosis and other diseases in descending order of the highest number of associated sentences.
Related references are mostly published in these journals:
| Disease | Cross reference | Weighted score | Related literature |
|---|
Possible diseases from mapped MeSH terms on references
We collected disease MeSH terms mapped to the references associated with PE(15:0/20:0)
PubChem Associated disorders and diseases
What pathways are associated with PE(15:0/20:0)
There are no associated biomedical information in the current reference collection.
PubChem Biomolecular Interactions and Pathways
Link to PubChem Biomolecular Interactions and PathwaysWhat cellular locations are associated with PE(15:0/20:0)?
Visualization in cellular structure
Associated locations are in red color. Not associated locations are in black.
Related references are published most in these journals:
| Location | Cross reference | Weighted score | Related literatures |
|---|
What functions are associated with PE(15:0/20:0)?
Related references are published most in these journals:
| Function | Cross reference | Weighted score | Related literatures |
|---|
What lipids are associated with PE(15:0/20:0)?
Related references are published most in these journals:
| Lipid concept | Cross reference | Weighted score | Related literatures |
|---|
What genes are associated with PE(15:0/20:0)?
Related references are published most in these journals:
| Gene | Cross reference | Weighted score | Related literatures |
|---|
What common seen animal models are associated with PE(15:0/20:0)?
Knock-out
Knock-out are used in the study 'Sequential synthesis and methylation of phosphatidylethanolamine promote lipid droplet biosynthesis and stability in tissue culture and in vivo.' (Hörl G et al., 2011) and Knock-out are used in the study 'An Atg4B mutant hampers the lipidation of LC3 paralogues and causes defects in autophagosome closure.' (Fujita N et al., 2008).
Cancer Model
Cancer Model are used in the study 'Improving penetration in tumors with nanoassemblies of phospholipids and doxorubicin.' (Tang N et al., 2007).
Related references are published most in these journals:
| Model | Cross reference | Weighted score | Related literatures |
|---|
NCBI Entrez Crosslinks
All references with PE(15:0/20:0)
Download all related citations| Authors | Title | Published | Journal | PubMed Link |
|---|---|---|---|---|
| Paran CW et al. | Lipogenesis mitigates dysregulated sarcoplasmic reticulum calcium uptake in muscular dystrophy. | 2015 | Biochim. Biophys. Acta | pmid:26361872 |
| Vrablik TL et al. | Lipidomic and proteomic analysis of Caenorhabditis elegans lipid droplets and identification of ACS-4 as a lipid droplet-associated protein. | 2015 | Biochim. Biophys. Acta | pmid:26121959 |
| Larson MC et al. | Phosphatidylethanolamine is externalized at the surface of microparticles. | 2012 | Biochim. Biophys. Acta | pmid:22960380 |
| Epand RF et al. | Bacterial lipid composition and the antimicrobial efficacy of cationic steroid compounds (Ceragenins). | 2007 | Biochim. Biophys. Acta | pmid:17599802 |
| Li Z et al. | A role for high density lipoproteins in hepatic phosphatidylcholine homeostasis. | 2007 | Biochim. Biophys. Acta | pmid:17513168 |
| Niebergall LJ and Vance DE | The ratio of phosphatidylcholine to phosphatidylethanolamine does not predict integrity of growing MT58 Chinese hamster ovary cells. | 2012 | Biochim. Biophys. Acta | pmid:22079326 |
| Di Bartolomeo F et al. | Involvement of a putative substrate binding site in the biogenesis and assembly of phosphatidylserine decarboxylase 1 from Saccharomyces cerevisiae. | 2017 | Biochim. Biophys. Acta | pmid:28473294 |
| Wang X et al. | Arginine-lysine positional swap of the LL-37 peptides reveals evolutional advantages of the native sequence and leads to bacterial probes. | 2017 | Biochim. Biophys. Acta | pmid:28450045 |
| Vieler A et al. | The influence of phase transitions in phosphatidylethanolamine models on the activity of violaxanthin de-epoxidase. | 2008 | Biochim. Biophys. Acta | pmid:18178148 |
| Guan Z et al. | The polar lipids of Clostridium psychrophilum, an anaerobic psychrophile. | 2013 | Biochim. Biophys. Acta | pmid:23454375 |
| Zhirnov AE et al. | Lipid composition determines interaction of liposome membranes with Pluronic L61. | 2005 | Biochim. Biophys. Acta | pmid:16405999 |
| Neumann J et al. | Diverse relations between ABC transporters and lipids: An overview. | 2017 | Biochim. Biophys. Acta | pmid:27693344 |
| Di Bartolomeo F et al. | Cell biology, physiology and enzymology of phosphatidylserine decarboxylase. | 2017 | Biochim. Biophys. Acta | pmid:27650064 |
| Willumeit R et al. | Structural rearrangement of model membranes by the peptide antibiotic NK-2. | 2005 | Biochim. Biophys. Acta | pmid:15893515 |
| Ruhanen H et al. | Depletion of TM6SF2 disturbs membrane lipid composition and dynamics in HuH7 hepatoma cells. | 2017 | Biochim. Biophys. Acta | pmid:28434889 |
| Goss R et al. | Lipid dependence of diadinoxanthin solubilization and de-epoxidation in artificial membrane systems resembling the lipid composition of the natural thylakoid membrane. | 2007 | Biochim. Biophys. Acta | pmid:16843433 |
| Tasseva G et al. | Lack of phosphatidylethanolamine N-methyltransferase in mice does not promote fatty acid oxidation in skeletal muscle. | 2016 | Biochim. Biophys. Acta | pmid:26603903 |
| Uyama T et al. | Characterization of the human tumor suppressors TIG3 and HRASLS2 as phospholipid-metabolizing enzymes. | 2009 | Biochim. Biophys. Acta | pmid:19615464 |
| Gabrys CM et al. | Nuclear magnetic resonance evidence for retention of a lamellar membrane phase with curvature in the presence of large quantities of the HIV fusion peptide. | 2010 | Biochim. Biophys. Acta | pmid:19616505 |
| Carmona-Antoñanzas G et al. | Molecular mechanism of dietary phospholipid requirement of Atlantic salmon, Salmo salar, fry. | 2015 | Biochim. Biophys. Acta | pmid:26303578 |