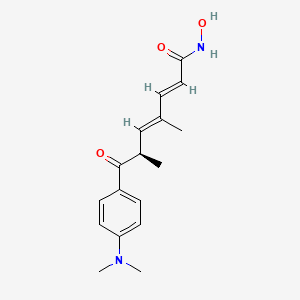
trichostatin A
Trichostatin is a lipid of Polyketides (PK) class. Trichostatin is associated with abnormalities such as Dentatorubral-Pallidoluysian Atrophy, PARAGANGLIOMAS 3, abnormal fragmented structure, Disintegration (morphologic abnormality) and Hyperostosis, Diffuse Idiopathic Skeletal. The involved functions are known as Acetylation, Cell Differentiation process, histone modification, Gene Silencing and Transcriptional Activation. Trichostatin often locates in CD41a, Hematopoietic System, Chromatin Structure, Blood and Endothelium. The associated genes with Trichostatin are SPI1 gene, CELL Gene, Chromatin, CXCR4 gene and DNMT1 gene. The related lipids are Butyrates, Promega, butyrate, Lipopolysaccharides and Steroids. The related experimental models are Knock-out, Mouse Model, Xenograft Model and Cancer Model.
Cross Reference
Introduction
To understand associated biological information of trichostatin A, we collected biological information of abnormalities, associated pathways, cellular/molecular locations, biological functions, related genes/proteins, lipids and common seen animal/experimental models with organized paragraphs from literatures.
What diseases are associated with trichostatin A?
trichostatin A is suspected in Infection, Morphologically altered structure, Ureteral obstruction, Photosensitization, Atherosclerosis, Hypertrophic Cardiomyopathy and other diseases in descending order of the highest number of associated sentences.
Related references are mostly published in these journals:
| Disease | Cross reference | Weighted score | Related literature |
|---|
Possible diseases from mapped MeSH terms on references
We collected disease MeSH terms mapped to the references associated with trichostatin A
PubChem Associated disorders and diseases
What pathways are associated with trichostatin A
Lipid pathways are not clear in current pathway databases. We organized associated pathways with trichostatin A through full-text articles, including metabolic pathways or pathways of biological mechanisms.
Related references are published most in these journals:
| Pathway name | Related literatures |
|---|
PubChem Biomolecular Interactions and Pathways
Link to PubChem Biomolecular Interactions and PathwaysWhat cellular locations are associated with trichostatin A?
Visualization in cellular structure
Associated locations are in red color. Not associated locations are in black.
Related references are published most in these journals:
| Location | Cross reference | Weighted score | Related literatures |
|---|
What functions are associated with trichostatin A?
Related references are published most in these journals:
| Function | Cross reference | Weighted score | Related literatures |
|---|
What lipids are associated with trichostatin A?
Related references are published most in these journals:
| Lipid concept | Cross reference | Weighted score | Related literatures |
|---|
What genes are associated with trichostatin A?
Related references are published most in these journals:
| Gene | Cross reference | Weighted score | Related literatures |
|---|
What common seen animal models are associated with trichostatin A?
Mouse Model
Mouse Model are used in the study 'Regulation of minichromosome maintenance gene family by microRNA-1296 and genistein in prostate cancer.' (Majid S et al., 2010), Mouse Model are used in the study 'Reversal of hypermethylation and reactivation of p16INK4a, RARbeta, and MGMT genes by genistein and other isoflavones from soy.' (Fang MZ et al., 2005) and Mouse Model are used in the study 'Histone deacetylase 3 mediates allergic skin inflammation by regulating expression of MCP1 protein.' (Kim Y et al., 2012).
Xenograft Model
Xenograft Model are used in the study 'Histone deacetylase inhibitors induce growth arrest and differentiation in uveal melanoma.' (Landreville S et al., 2012), Xenograft Model are used in the study 'Extended treatment with physiologic concentrations of dietary phytochemicals results in altered gene expression, reduced growth, and apoptosis of cancer cells.' (Moiseeva EP et al., 2007) and Xenograft Model are used in the study 'Retinoic acid and the histone deacetylase inhibitor trichostatin a inhibit the proliferation of human renal cell carcinoma in a xenograft tumor model.' (Touma SE et al., 2005).
Cancer Model
Cancer Model are used in the study 'Plasma pharmacokinetics and metabolism of the histone deacetylase inhibitor trichostatin a after intraperitoneal administration to mice.' (Sanderson L et al., 2004).
Related references are published most in these journals:
| Model | Cross reference | Weighted score | Related literatures |
|---|
NCBI Entrez Crosslinks
All references with trichostatin A
Download all related citations| Authors | Title | Published | Journal | PubMed Link |
|---|---|---|---|---|
| Henderson C et al. | Role of caspases, Bid, and p53 in the apoptotic response triggered by histone deacetylase inhibitors trichostatin-A (TSA) and suberoylanilide hydroxamic acid (SAHA). | 2003 | J. Biol. Chem. | pmid:12556448 |
| Tokési N et al. | TPPP/p25 promotes tubulin acetylation by inhibiting histone deacetylase 6. | 2010 | J. Biol. Chem. | pmid:20308065 |
| Varshochi R et al. | ICI182,780 induces p21Waf1 gene transcription through releasing histone deacetylase 1 and estrogen receptor alpha from Sp1 sites to induce cell cycle arrest in MCF-7 breast cancer cell line. | 2005 | J. Biol. Chem. | pmid:15557281 |
| Corvetta D et al. | Physical interaction between MYCN oncogene and polycomb repressive complex 2 (PRC2) in neuroblastoma: functional and therapeutic implications. | 2013 | J. Biol. Chem. | pmid:23362253 |
| Liou JS et al. | Oncogenic ras mediates apoptosis in response to protein kinase C inhibition through the generation of reactive oxygen species. | 2000 | J. Biol. Chem. | pmid:10967125 |
| Wee G et al. | Inheritable histone H4 acetylation of somatic chromatins in cloned embryos. | 2006 | J. Biol. Chem. | pmid:16371357 |
| Stimson KM and Vertino PM | Methylation-mediated silencing of TMS1/ASC is accompanied by histone hypoacetylation and CpG island-localized changes in chromatin architecture. | 2002 | J. Biol. Chem. | pmid:11733524 |
| Hsu MC et al. | HER-2/neu represses the metastasis suppressor RECK via ERK and Sp transcription factors to promote cell invasion. | 2006 | J. Biol. Chem. | pmid:16377629 |
| Yang J et al. | A novel mechanism involving coordinated regulation of nuclear levels and acetylation of NF-YA and Bcl6 activates RGS4 transcription. | 2010 | J. Biol. Chem. | pmid:20630860 |
| Kurschat P et al. | Neuron restrictive silencer factor NRSF/REST is a transcriptional repressor of neuropilin-1 and diminishes the ability of semaphorin 3A to inhibit keratinocyte migration. | 2006 | J. Biol. Chem. | pmid:16330548 |
| Ooi L et al. | BRG1 chromatin remodeling activity is required for efficient chromatin binding by repressor element 1-silencing transcription factor (REST) and facilitates REST-mediated repression. | 2006 | J. Biol. Chem. | pmid:17023429 |
| Topark-Ngarm A et al. | CTIP2 associates with the NuRD complex on the promoter of p57KIP2, a newly identified CTIP2 target gene. | 2006 | J. Biol. Chem. | pmid:16950772 |
| Kurtev V et al. | Transcriptional regulation by the repressor of estrogen receptor activity via recruitment of histone deacetylases. | 2004 | J. Biol. Chem. | pmid:15140878 |
| Sekimata M et al. | Involvement of a novel zinc finger protein, MIZF, in transcriptional repression by interacting with a methyl-CpG-binding protein, MBD2. | 2001 | J. Biol. Chem. | pmid:11553631 |
| Roy S and Tenniswood M | Site-specific acetylation of p53 directs selective transcription complex assembly. | 2007 | J. Biol. Chem. | pmid:17121856 |
| Butler PL et al. | Acetylation stimulates the epithelial sodium channel by reducing its ubiquitination and degradation. | 2015 | J. Biol. Chem. | pmid:25787079 |
| Kuno A et al. | Resveratrol improves cardiomyopathy in dystrophin-deficient mice through SIRT1 protein-mediated modulation of p300 protein. | 2013 | J. Biol. Chem. | pmid:23297412 |
| Musikacharoen T et al. | Histone acetylation and activation of cAMP-response element-binding protein regulate transcriptional activation of MKP-M in lipopolysaccharide-stimulated macrophages. | 2003 | J. Biol. Chem. | pmid:12511574 |
| Gusterson RJ et al. | The transcriptional co-activators CREB-binding protein (CBP) and p300 play a critical role in cardiac hypertrophy that is dependent on their histone acetyltransferase activity. | 2003 | J. Biol. Chem. | pmid:12477714 |
| Phiel CJ et al. | Histone deacetylase is a direct target of valproic acid, a potent anticonvulsant, mood stabilizer, and teratogen. | 2001 | J. Biol. Chem. | pmid:11473107 |
| Dai X et al. | Somatic nucleus reprogramming is significantly improved by m-carboxycinnamic acid bishydroxamide, a histone deacetylase inhibitor. | 2010 | J. Biol. Chem. | pmid:20566633 |
| Leu TH et al. | Participation of p97Eps8 in Src-mediated transformation. | 2004 | J. Biol. Chem. | pmid:14699156 |
| Zhang L et al. | Inhibition of histone deacetylase-induced myocardial repair is mediated by c-kit in infarcted hearts. | 2012 | J. Biol. Chem. | pmid:23024362 |
| Pagans S et al. | Repression by TTK69 of GAGA-mediated activation occurs in the absence of TTK69 binding to DNA and solely requires the contribution of the POZ/BTB domain of TTK69. | 2004 | J. Biol. Chem. | pmid:14701830 |
| Hwang CK et al. | Transcriptional regulation of mouse mu opioid receptor gene by PU.1. | 2004 | J. Biol. Chem. | pmid:14998994 |
| Brush MH et al. | Deactylase inhibitors disrupt cellular complexes containing protein phosphatases and deacetylases. | 2004 | J. Biol. Chem. | pmid:14670976 |
| Qiao Y et al. | Dual roles of histone H3 lysine 9 acetylation in human embryonic stem cell pluripotency and neural differentiation. | 2015 | J. Biol. Chem. | pmid:25519907 |
| Hattori N et al. | Epigenetic control of mouse Oct-4 gene expression in embryonic stem cells and trophoblast stem cells. | 2004 | J. Biol. Chem. | pmid:14761969 |
| Horion J et al. | Histone deacetylase inhibitor trichostatin A sustains sodium pervanadate-induced NF-kappaB activation by delaying IkappaBalpha mRNA resynthesis: comparison with tumor necrosis factor alpha. | 2007 | J. Biol. Chem. | pmid:17409387 |
| Kiefer SM et al. | Murine Sall1 represses transcription by recruiting a histone deacetylase complex. | 2002 | J. Biol. Chem. | pmid:11836251 |
| Taniura S et al. | Transcriptional regulation of cyclooxygenase-1 by histone deacetylase inhibitors in normal human astrocyte cells. | 2002 | J. Biol. Chem. | pmid:11877441 |
| Senawong T et al. | Involvement of the histone deacetylase SIRT1 in chicken ovalbumin upstream promoter transcription factor (COUP-TF)-interacting protein 2-mediated transcriptional repression. | 2003 | J. Biol. Chem. | pmid:12930829 |
| Ginjala V et al. | Multiple cis elements within the Igf2/H19 insulator domain organize a distance-dependent silencer. A cautionary note. | 2002 | J. Biol. Chem. | pmid:11777900 |
| Massillon D et al. | Regulation of glucose-6-phosphatase gene expression in cultured hepatocytes and H4IIE cells by short-chain fatty acids: role of hepatic nuclear factor-4alpha. | 2003 | J. Biol. Chem. | pmid:12915406 |
| Cabral AL et al. | Regulation of the cellular prion protein gene expression depends on chromatin conformation. | 2002 | J. Biol. Chem. | pmid:11739375 |
| Park SH et al. | Transcriptional regulation of the transforming growth factor beta type II receptor gene by histone acetyltransferase and deacetylase is mediated by NF-Y in human breast cancer cells. | 2002 | J. Biol. Chem. | pmid:11744689 |
| Wang Y et al. | Nuclear factor κB mediates suppression of canonical transient receptor potential 6 expression by reactive oxygen species and protein kinase C in kidney cells. | 2013 | J. Biol. Chem. | pmid:23525112 |
| Singal R et al. | Methylation of promoter proximal-transcribed sequences of an embryonic globin gene inhibits transcription in primary erythroid cells and promotes formation of a cell type-specific methyl cytosine binding complex. | 2002 | J. Biol. Chem. | pmid:11684679 |
| Nam HJ et al. | Clostridium difficile toxin A decreases acetylation of tubulin, leading to microtubule depolymerization through activation of histone deacetylase 6, and this mediates acute inflammation. | 2010 | J. Biol. Chem. | pmid:20696758 |
| Ammanamanchi S et al. | Acetylated sp3 is a transcriptional activator. | 2003 | J. Biol. Chem. | pmid:12837748 |
| Cribbs A et al. | Inhibition of histone H3K27 demethylases selectively modulates inflammatory phenotypes of natural killer cells. | 2018 | J. Biol. Chem. | pmid:29301935 |
| Sarg B et al. | Histone H4 hyperacetylation precludes histone H4 lysine 20 trimethylation. | 2004 | J. Biol. Chem. | pmid:15456746 |
| Yoshida T et al. | Kruppel-like factor 4 protein regulates isoproterenol-induced cardiac hypertrophy by modulating myocardin expression and activity. | 2014 | J. Biol. Chem. | pmid:25100730 |
| Lee SK et al. | Silencing mediator of retinoic acid and thyroid hormone receptors, as a novel transcriptional corepressor molecule of activating protein-1, nuclear factor-kappaB, and serum response factor. | 2000 | J. Biol. Chem. | pmid:10777532 |
| Fu M et al. | p300 and p300/cAMP-response element-binding protein-associated factor acetylate the androgen receptor at sites governing hormone-dependent transactivation. | 2000 | J. Biol. Chem. | pmid:10779504 |
| Yoshida M et al. | Potent and specific inhibition of mammalian histone deacetylase both in vivo and in vitro by trichostatin A. | 1990 | J. Biol. Chem. | pmid:2211619 |
| Lauffer BE et al. | Histone deacetylase (HDAC) inhibitor kinetic rate constants correlate with cellular histone acetylation but not transcription and cell viability. | 2013 | J. Biol. Chem. | pmid:23897821 |
| Diamond SE and Gutierrez-Hartmann A | The Pit-1beta domain dictates active repression and alteration of histone acetylation of the proximal prolactin promoter. | 2000 | J. Biol. Chem. | pmid:10921928 |
| Spallotta F et al. | A nitric oxide-dependent cross-talk between class I and III histone deacetylases accelerates skin repair. | 2013 | J. Biol. Chem. | pmid:23463510 |
| Di-Poï N et al. | Transcriptional repression of peroxisome proliferator-activated receptor beta/delta in murine keratinocytes by CCAAT/enhancer-binding proteins. | 2005 | J. Biol. Chem. | pmid:16166081 |
| Mogal AP et al. | Haploinsufficient prostate tumor suppression by Nkx3.1: a role for chromatin accessibility in dosage-sensitive gene regulation. | 2007 | J. Biol. Chem. | pmid:17602165 |
| Guo Y et al. | Regulation of DNA methylation in human breast cancer. Effect on the urokinase-type plasminogen activator gene production and tumor invasion. | 2002 | J. Biol. Chem. | pmid:12198113 |
| Wong CY et al. | Epigenetic regulation of phosphatidylinositol 3,4,5-triphosphate-dependent Rac exchanger 1 gene expression in prostate cancer cells. | 2011 | J. Biol. Chem. | pmid:21636851 |
| Chen CS et al. | Histone acetylation-independent effect of histone deacetylase inhibitors on Akt through the reshuffling of protein phosphatase 1 complexes. | 2005 | J. Biol. Chem. | pmid:16186112 |
| Zschocke J et al. | Estrogen receptor alpha-mediated silencing of caveolin gene expression in neuronal cells. | 2002 | J. Biol. Chem. | pmid:12138116 |
| Zhou Q et al. | Rapid induction of histone hyperacetylation and cellular differentiation in human breast tumor cell lines following degradation of histone deacetylase-1. | 2000 | J. Biol. Chem. | pmid:10938272 |
| Ventura A et al. | The p66Shc longevity gene is silenced through epigenetic modifications of an alternative promoter. | 2002 | J. Biol. Chem. | pmid:11948181 |
| Kadiyala V et al. | Class I lysine deacetylases facilitate glucocorticoid-induced transcription. | 2013 | J. Biol. Chem. | pmid:23946490 |
| Gupta MP et al. | HDAC4 and PCAF bind to cardiac sarcomeres and play a role in regulating myofilament contractile activity. | 2008 | J. Biol. Chem. | pmid:18250163 |
| Milutinovic S et al. | Proliferating cell nuclear antigen associates with histone deacetylase activity, integrating DNA replication and chromatin modification. | 2002 | J. Biol. Chem. | pmid:11929879 |
| Kang J et al. | Heat shock protein 90 mediates protein-protein interactions between human aminoacyl-tRNA synthetases. | 2000 | J. Biol. Chem. | pmid:10913161 |
| Qian Y et al. | DeltaNp63, a target of DEC1 and histone deacetylase 2, modulates the efficacy of histone deacetylase inhibitors in growth suppression and keratinocyte differentiation. | 2011 | J. Biol. Chem. | pmid:21317427 |
| Huang H et al. | MEIS C termini harbor transcriptional activation domains that respond to cell signaling. | 2005 | J. Biol. Chem. | pmid:15654074 |
| Shimazu T et al. | Multiple histone deacetylases and the CREB-binding protein regulate pre-mRNA 3'-end processing. | 2007 | J. Biol. Chem. | pmid:17172643 |
| Ohoka N et al. | Critical and functional regulation of CHOP (C/EBP homologous protein) through the N-terminal portion. | 2007 | J. Biol. Chem. | pmid:17872950 |
| Tong X et al. | Cyclooxygenase-2 regulation in colon cancer cells: modulation of RNA polymerase II elongation by histone deacetylase inhibitors. | 2005 | J. Biol. Chem. | pmid:15713675 |
| Gan Y et al. | Role of histone deacetylation in cell-specific expression of endothelial nitric-oxide synthase. | 2005 | J. Biol. Chem. | pmid:15722551 |
| McCool KW et al. | The role of histone acetylation in regulating early gene expression patterns during early embryonic stem cell differentiation. | 2007 | J. Biol. Chem. | pmid:17204470 |
| Wen J et al. | Tal1/SCL binding to pericentromeric DNA represses transcription. | 2005 | J. Biol. Chem. | pmid:15677454 |
| Zhang Y and Dufau ML | Silencing of transcription of the human luteinizing hormone receptor gene by histone deacetylase-mSin3A complex. | 2002 | J. Biol. Chem. | pmid:12091390 |
| Lin HM et al. | Cyclin D1 Is a Ligand-independent Co-repressor for Thyroid Hormone Receptors. | 2002 | J. Biol. Chem. | pmid:12048199 |
| Gong XQ and Li L | Dermo-1, a multifunctional basic helix-loop-helix protein, represses MyoD transactivation via the HLH domain, MEF2 interaction, and chromatin deacetylation. | 2002 | J. Biol. Chem. | pmid:11809751 |
| Kizuka Y et al. | Epigenetic regulation of a brain-specific glycosyltransferase N-acetylglucosaminyltransferase-IX (GnT-IX) by specific chromatin modifiers. | 2014 | J. Biol. Chem. | pmid:24619417 |
| Chang I and Wang CY | Inhibition of HDAC6 Protein Enhances Bortezomib-induced Apoptosis in Head and Neck Squamous Cell Carcinoma (HNSCC) by Reducing Autophagy. | 2016 | J. Biol. Chem. | pmid:27369083 |
| Kumar P et al. | Histone deacetylase inhibitors modulate the transcriptional regulation of guanylyl cyclase/natriuretic peptide receptor-a gene: interactive roles of modified histones, histone acetyltransferase, p300, AND Sp1. | 2014 | J. Biol. Chem. | pmid:24451378 |
| Verdel A and Khochbin S | Identification of a new family of higher eukaryotic histone deacetylases. Coordinate expression of differentiation-dependent chromatin modifiers. | 1999 | J. Biol. Chem. | pmid:9891014 |
| Tominaga K et al. | MRGX is a novel transcriptional regulator that exhibits activation or repression of the B-myb promoter in a cell type-dependent manner. | 2003 | J. Biol. Chem. | pmid:14506250 |
| Schuetz A et al. | Human HDAC7 harbors a class IIa histone deacetylase-specific zinc binding motif and cryptic deacetylase activity. | 2008 | J. Biol. Chem. | pmid:18285338 |
| Gradin K et al. | Repression of dioxin signal transduction in fibroblasts. Identification Of a putative repressor associated with Arnt. | 1999 | J. Biol. Chem. | pmid:10224119 |
| Zhang Y et al. | Unlocking repression of the human luteinizing hormone receptor gene by trichostatin A-induced cell-specific phosphatase release. | 2008 | J. Biol. Chem. | pmid:18596044 |
| Agoulnik IU et al. | Repressors of androgen and progesterone receptor action. | 2003 | J. Biol. Chem. | pmid:12771131 |
| Yamaguchi K et al. | Histone deacetylase inhibitors suppress the induction of c-Jun and its target genes including COX-2. | 2005 | J. Biol. Chem. | pmid:15994313 |
| Kramps C et al. | E2F and Sp1/Sp3 Synergize but are not sufficient to activate the MYCN gene in neuroblastomas. | 2004 | J. Biol. Chem. | pmid:14645238 |
| Marks J et al. | Two functionally divergent p53-responsive elements in the rat bradykinin B2 receptor promoter. | 2003 | J. Biol. Chem. | pmid:12791684 |
| Vanden Berghe W et al. | The nuclear factor-kappaB engages CBP/p300 and histone acetyltransferase activity for transcriptional activation of the interleukin-6 gene promoter. | 1999 | J. Biol. Chem. | pmid:10542243 |
| Gray SG et al. | Functional characterization of JMJD2A, a histone deacetylase- and retinoblastoma-binding protein. | 2005 | J. Biol. Chem. | pmid:15927959 |
| Asthana J et al. | Inhibition of HDAC6 deacetylase activity increases its binding with microtubules and suppresses microtubule dynamic instability in MCF-7 cells. | 2013 | J. Biol. Chem. | pmid:23798680 |
| Wagner S et al. | SET-mediated promoter hypoacetylation is a prerequisite for coactivation of the estrogen-responsive pS2 gene by PRMT1. | 2006 | J. Biol. Chem. | pmid:16861234 |
| Tagami T et al. | Mechanisms that mediate negative regulation of the thyroid-stimulating hormone alpha gene by the thyroid hormone receptor. | 1999 | J. Biol. Chem. | pmid:10428804 |
| Hayakawa F et al. | Acetylation of PML is involved in histone deacetylase inhibitor-mediated apoptosis. | 2008 | J. Biol. Chem. | pmid:18621739 |
| Fass DM et al. | Deacetylase activity is required for cAMP activation of a subset of CREB target genes. | 2003 | J. Biol. Chem. | pmid:12939274 |
| Zhao C et al. | A composite motif of the Drosophila morphogenetic protein bicoid critical to transcription control. | 2003 | J. Biol. Chem. | pmid:12939280 |
| Hinoi T et al. | Silencing of CDX2 expression in colon cancer via a dominant repression pathway. | 2003 | J. Biol. Chem. | pmid:12947088 |
| Hichino A et al. | Down-regulation of Claudin-2 Expression and Proliferation by Epigenetic Inhibitors in Human Lung Adenocarcinoma A549 Cells. | 2017 | J. Biol. Chem. | pmid:28057758 |
| Pulukuri SM et al. | Inhibition of histone deacetylase activity promotes invasion of human cancer cells through activation of urokinase plasminogen activator. | 2007 | J. Biol. Chem. | pmid:17923479 |
| Webster SJ et al. | Regulation of GTP-binding protein (Gαs) expression in human myometrial cells: a role for tumor necrosis factor in modulating Gαs promoter acetylation by transcriptional complexes. | 2013 | J. Biol. Chem. | pmid:23297421 |
| Frescas D et al. | Nuclear trapping of the forkhead transcription factor FoxO1 via Sirt-dependent deacetylation promotes expression of glucogenetic genes. | 2005 | J. Biol. Chem. | pmid:15788402 |
| Levenson JM et al. | Regulation of histone acetylation during memory formation in the hippocampus. | 2004 | J. Biol. Chem. | pmid:15273246 |
| Banchio C et al. | Role of histone deacetylase in the expression of CTP:phosphocholine cytidylyltransferase alpha. | 2006 | J. Biol. Chem. | pmid:16484221 |
| Arora T et al. | PIASx is a transcriptional co-repressor of signal transducer and activator of transcription 4. | 2003 | J. Biol. Chem. | pmid:12716907 |