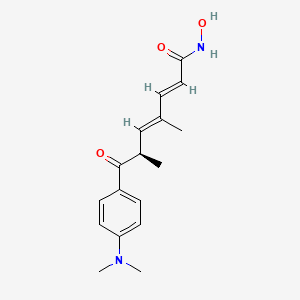
trichostatin A
Trichostatin is a lipid of Polyketides (PK) class. Trichostatin is associated with abnormalities such as Dentatorubral-Pallidoluysian Atrophy, PARAGANGLIOMAS 3, abnormal fragmented structure, Disintegration (morphologic abnormality) and Hyperostosis, Diffuse Idiopathic Skeletal. The involved functions are known as Acetylation, Cell Differentiation process, histone modification, Gene Silencing and Transcriptional Activation. Trichostatin often locates in CD41a, Hematopoietic System, Chromatin Structure, Blood and Endothelium. The associated genes with Trichostatin are SPI1 gene, CELL Gene, Chromatin, CXCR4 gene and DNMT1 gene. The related lipids are Butyrates, Promega, butyrate, Lipopolysaccharides and Steroids. The related experimental models are Knock-out, Mouse Model, Xenograft Model and Cancer Model.
Cross Reference
Introduction
To understand associated biological information of trichostatin A, we collected biological information of abnormalities, associated pathways, cellular/molecular locations, biological functions, related genes/proteins, lipids and common seen animal/experimental models with organized paragraphs from literatures.
What diseases are associated with trichostatin A?
trichostatin A is suspected in Infection, Morphologically altered structure, Ureteral obstruction, Photosensitization, Atherosclerosis, Hypertrophic Cardiomyopathy and other diseases in descending order of the highest number of associated sentences.
Related references are mostly published in these journals:
| Disease | Cross reference | Weighted score | Related literature |
|---|
Possible diseases from mapped MeSH terms on references
We collected disease MeSH terms mapped to the references associated with trichostatin A
PubChem Associated disorders and diseases
What pathways are associated with trichostatin A
Lipid pathways are not clear in current pathway databases. We organized associated pathways with trichostatin A through full-text articles, including metabolic pathways or pathways of biological mechanisms.
Related references are published most in these journals:
| Pathway name | Related literatures |
|---|
PubChem Biomolecular Interactions and Pathways
Link to PubChem Biomolecular Interactions and PathwaysWhat cellular locations are associated with trichostatin A?
Visualization in cellular structure
Associated locations are in red color. Not associated locations are in black.
Related references are published most in these journals:
| Location | Cross reference | Weighted score | Related literatures |
|---|
What functions are associated with trichostatin A?
Related references are published most in these journals:
| Function | Cross reference | Weighted score | Related literatures |
|---|
What lipids are associated with trichostatin A?
Related references are published most in these journals:
| Lipid concept | Cross reference | Weighted score | Related literatures |
|---|
What genes are associated with trichostatin A?
Related references are published most in these journals:
| Gene | Cross reference | Weighted score | Related literatures |
|---|
What common seen animal models are associated with trichostatin A?
Mouse Model
Mouse Model are used in the study 'Regulation of minichromosome maintenance gene family by microRNA-1296 and genistein in prostate cancer.' (Majid S et al., 2010), Mouse Model are used in the study 'Reversal of hypermethylation and reactivation of p16INK4a, RARbeta, and MGMT genes by genistein and other isoflavones from soy.' (Fang MZ et al., 2005) and Mouse Model are used in the study 'Histone deacetylase 3 mediates allergic skin inflammation by regulating expression of MCP1 protein.' (Kim Y et al., 2012).
Xenograft Model
Xenograft Model are used in the study 'Histone deacetylase inhibitors induce growth arrest and differentiation in uveal melanoma.' (Landreville S et al., 2012), Xenograft Model are used in the study 'Extended treatment with physiologic concentrations of dietary phytochemicals results in altered gene expression, reduced growth, and apoptosis of cancer cells.' (Moiseeva EP et al., 2007) and Xenograft Model are used in the study 'Retinoic acid and the histone deacetylase inhibitor trichostatin a inhibit the proliferation of human renal cell carcinoma in a xenograft tumor model.' (Touma SE et al., 2005).
Cancer Model
Cancer Model are used in the study 'Plasma pharmacokinetics and metabolism of the histone deacetylase inhibitor trichostatin a after intraperitoneal administration to mice.' (Sanderson L et al., 2004).
Related references are published most in these journals:
| Model | Cross reference | Weighted score | Related literatures |
|---|
NCBI Entrez Crosslinks
All references with trichostatin A
Download all related citations| Authors | Title | Published | Journal | PubMed Link |
|---|---|---|---|---|
| Im H et al. | Histone deacetylase-dependent establishment and maintenance of broad low-level histone acetylation within a tissue-specific chromatin domain. | 2002 | Biochemistry | pmid:12484752 |
| Darlyuk-Saadon I et al. | The bZIP repressor proteins, c-Jun dimerization protein 2 and activating transcription factor 3, recruit multiple HDAC members to the ATF3 promoter. | 2012 Nov-Dec | Biochim. Biophys. Acta | pmid:22989952 |
| Lugrin J et al. | Histone deacetylase inhibitors repress macrophage migration inhibitory factor (MIF) expression by targeting MIF gene transcription through a local chromatin deacetylation. | 2009 | Biochim. Biophys. Acta | pmid:19747950 |
| Cheli Y et al. | Transcriptional and epigenetic regulation of the integrin collagen receptor locus ITGA1-PELO-ITGA2. | 2007 Sep-Oct | Biochim. Biophys. Acta | pmid:17669516 |
| Lee KJ et al. | Downregulation of a tumor suppressor RECK by hypoxia through recruitment of HDAC1 and HIF-1alpha to reverse HRE site in the promoter. | 2010 | Biochim. Biophys. Acta | pmid:20080132 |
| Barrasa JI et al. | Histone deacetylase inhibitors upregulate MMP11 gene expression through Sp1/Smad complexes in human colon adenocarcinoma cells. | 2012 | Biochim. Biophys. Acta | pmid:22227581 |
| Hsu YF et al. | Trichostatin A and sirtinol suppressed survivin expression through AMPK and p38MAPK in HT29 colon cancer cells. | 2012 | Biochim. Biophys. Acta | pmid:22155142 |
| Zelivianski S et al. | Multipathways for transdifferentiation of human prostate cancer cells into neuroendocrine-like phenotype. | 2001 | Biochim. Biophys. Acta | pmid:11389966 |
| Hsu YF et al. | p53 in trichostatin A induced C6 glioma cell death. | 2011 | Biochim. Biophys. Acta | pmid:21376104 |
| Laribee RN and Klemsz MJ | Histone H4 HDAC activity is necessary for expression of the PU.1 gene. | 2005 | Biochim. Biophys. Acta | pmid:16139904 |
| Chen W et al. | MicroRNA-455-3p modulates cartilage development and degeneration through modification of histone H3 acetylation. | 2016 | Biochim. Biophys. Acta | pmid:27638301 |
| Alcarraz-Vizán G et al. | Validation of NCM460 cell model as control in antitumor strategies targeting colon adenocarcinoma metabolic reprogramming: trichostatin A as a case study. | 2014 | Biochim. Biophys. Acta | pmid:24368265 |
| Hsu YF et al. | MAPK phosphatase-1 contributes to trichostatin A inhibition of cyclooxygenase-2 expression in human umbilical vascular endothelial cells exposed to lipopolysaccharide. | 2011 | Biochim. Biophys. Acta | pmid:21911040 |
| Sharma K et al. | Constitutive hyperactivity of histone deacetylases enhances radioresistance in Lepidopteran Sf9 insect cells. | 2016 | Biochim. Biophys. Acta | pmid:26968462 |
| Cui W et al. | A SUMO-acetyl switch in PXR biology. | 2016 | Biochim. Biophys. Acta | pmid:26883953 |
| Gobl AE et al. | Menin represses JunD-activated transcription by a histone deacetylase-dependent mechanism. | 1999 | Biochim. Biophys. Acta | pmid:10500243 |
| Stoppani E et al. | Point mutated caveolin-3 form (P104L) impairs myoblast differentiation via Akt and p38 signalling reduction, leading to an immature cell signature. | 2011 | Biochim. Biophys. Acta | pmid:21182936 |
| Moore PS et al. | Gene expression profiling after treatment with the histone deacetylase inhibitor trichostatin A reveals altered expression of both pro- and anti-apoptotic genes in pancreatic adenocarcinoma cells. | 2004 | Biochim. Biophys. Acta | pmid:15363630 |
| Schnur N et al. | The histone deacetylase inhibitor trichostatin A mediates upregulation of 5-lipoxygenase promoter activity by recruitment of Sp1 to distinct GC-boxes. | 2007 | Biochim. Biophys. Acta | pmid:17894944 |
| Putnik J et al. | The interaction of ETV6 (TEL) and TIP60 requires a functional histone acetyltransferase domain in TIP60. | 2007 | Biochim. Biophys. Acta | pmid:17980166 |