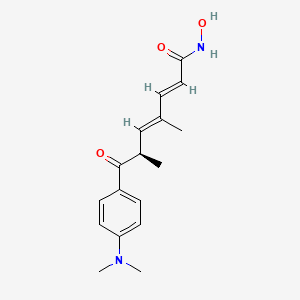
trichostatin A
Trichostatin is a lipid of Polyketides (PK) class. Trichostatin is associated with abnormalities such as Dentatorubral-Pallidoluysian Atrophy, PARAGANGLIOMAS 3, abnormal fragmented structure, Disintegration (morphologic abnormality) and Hyperostosis, Diffuse Idiopathic Skeletal. The involved functions are known as Acetylation, Cell Differentiation process, histone modification, Gene Silencing and Transcriptional Activation. Trichostatin often locates in CD41a, Hematopoietic System, Chromatin Structure, Blood and Endothelium. The associated genes with Trichostatin are SPI1 gene, CELL Gene, Chromatin, CXCR4 gene and DNMT1 gene. The related lipids are Butyrates, Promega, butyrate, Lipopolysaccharides and Steroids. The related experimental models are Knock-out, Mouse Model, Xenograft Model and Cancer Model.
Cross Reference
Introduction
To understand associated biological information of trichostatin A, we collected biological information of abnormalities, associated pathways, cellular/molecular locations, biological functions, related genes/proteins, lipids and common seen animal/experimental models with organized paragraphs from literatures.
What diseases are associated with trichostatin A?
trichostatin A is suspected in Infection, Morphologically altered structure, Ureteral obstruction, Photosensitization, Atherosclerosis, Hypertrophic Cardiomyopathy and other diseases in descending order of the highest number of associated sentences.
Related references are mostly published in these journals:
| Disease | Cross reference | Weighted score | Related literature |
|---|
Possible diseases from mapped MeSH terms on references
We collected disease MeSH terms mapped to the references associated with trichostatin A
PubChem Associated disorders and diseases
What pathways are associated with trichostatin A
Lipid pathways are not clear in current pathway databases. We organized associated pathways with trichostatin A through full-text articles, including metabolic pathways or pathways of biological mechanisms.
Related references are published most in these journals:
| Pathway name | Related literatures |
|---|
PubChem Biomolecular Interactions and Pathways
Link to PubChem Biomolecular Interactions and PathwaysWhat cellular locations are associated with trichostatin A?
Visualization in cellular structure
Associated locations are in red color. Not associated locations are in black.
Related references are published most in these journals:
| Location | Cross reference | Weighted score | Related literatures |
|---|
What functions are associated with trichostatin A?
Related references are published most in these journals:
| Function | Cross reference | Weighted score | Related literatures |
|---|
What lipids are associated with trichostatin A?
Related references are published most in these journals:
| Lipid concept | Cross reference | Weighted score | Related literatures |
|---|
What genes are associated with trichostatin A?
Related references are published most in these journals:
| Gene | Cross reference | Weighted score | Related literatures |
|---|
What common seen animal models are associated with trichostatin A?
Mouse Model
Mouse Model are used in the study 'Regulation of minichromosome maintenance gene family by microRNA-1296 and genistein in prostate cancer.' (Majid S et al., 2010), Mouse Model are used in the study 'Reversal of hypermethylation and reactivation of p16INK4a, RARbeta, and MGMT genes by genistein and other isoflavones from soy.' (Fang MZ et al., 2005) and Mouse Model are used in the study 'Histone deacetylase 3 mediates allergic skin inflammation by regulating expression of MCP1 protein.' (Kim Y et al., 2012).
Xenograft Model
Xenograft Model are used in the study 'Histone deacetylase inhibitors induce growth arrest and differentiation in uveal melanoma.' (Landreville S et al., 2012), Xenograft Model are used in the study 'Extended treatment with physiologic concentrations of dietary phytochemicals results in altered gene expression, reduced growth, and apoptosis of cancer cells.' (Moiseeva EP et al., 2007) and Xenograft Model are used in the study 'Retinoic acid and the histone deacetylase inhibitor trichostatin a inhibit the proliferation of human renal cell carcinoma in a xenograft tumor model.' (Touma SE et al., 2005).
Cancer Model
Cancer Model are used in the study 'Plasma pharmacokinetics and metabolism of the histone deacetylase inhibitor trichostatin a after intraperitoneal administration to mice.' (Sanderson L et al., 2004).
Related references are published most in these journals:
| Model | Cross reference | Weighted score | Related literatures |
|---|
NCBI Entrez Crosslinks
All references with trichostatin A
Download all related citations| Authors | Title | Published | Journal | PubMed Link |
|---|---|---|---|---|
| Wang LG et al. | Androgen receptor level controlled by a suppressor complex lost in an androgen-independent prostate cancer cell line. | 2004 | Oncogene | pmid:15156193 |
| Yokota T et al. | Histone deacetylase inhibitors activate INK4d gene through Sp1 site in its promoter. | 2004 | Oncogene | pmid:15107822 |
| Suzuki M et al. | Site-specific DNA methylation by a complex of PU.1 and Dnmt3a/b. | 2006 | Oncogene | pmid:16331260 |
| McGarry LC et al. | Invasion of v-Fos(FBR)-transformed cells is dependent upon histone deacetylase activity and suppression of histone deacetylase regulated genes. | 2004 | Oncogene | pmid:15107823 |
| Guo WH et al. | Inhibition of growth of mouse gastric cancer cells by Runx3, a novel tumor suppressor. | 2002 | Oncogene | pmid:12447699 |
| Cao K et al. | Histone deacetylase inhibitors prevent activation-induced cell death and promote anti-tumor immunity. | 2015 | Oncogene | pmid:25745993 |
| Ong DC et al. | LARG at chromosome 11q23 has functional characteristics of a tumor suppressor in human breast and colorectal cancer. | 2009 | Oncogene | pmid:19734946 |
| Tessema M et al. | Re-expression of CXCL14, a common target for epigenetic silencing in lung cancer, induces tumor necrosis. | 2010 | Oncogene | pmid:20562917 |
| Vincent A et al. | Epigenetic regulation (DNA methylation, histone modifications) of the 11p15 mucin genes (MUC2, MUC5AC, MUC5B, MUC6) in epithelial cancer cells. | 2007 | Oncogene | pmid:17471237 |
| Konduri SD et al. | Promoter methylation and silencing of the tissue factor pathway inhibitor-2 (TFPI-2), a gene encoding an inhibitor of matrix metalloproteinases in human glioma cells. | 2003 | Oncogene | pmid:12881707 |
| Mulholland NM et al. | Inhibition of MMTV transcription by HDAC inhibitors occurs independent of changes in chromatin remodeling and increased histone acetylation. | 2003 | Oncogene | pmid:12894222 |
| Wong CH et al. | CTSL2 is a pro-apoptotic target of E2F1 and a modulator of histone deacetylase inhibitor and DNA damage-induced apoptosis. | 2014 | Oncogene | pmid:23542171 |
| Augenlicht L et al. | Repression of MUC2 gene expression by butyrate, a physiological regulator of intestinal cell maturation. | 2003 | Oncogene | pmid:12902981 |
| Gurova KV et al. | Expression of prostate specific antigen (PSA) is negatively regulated by p53. | 2002 | Oncogene | pmid:11791186 |
| Zhong S et al. | Phosphorylation at serine 28 and acetylation at lysine 9 of histone H3 induced by trichostatin A. | 2003 | Oncogene | pmid:12917630 |
| Quan H et al. | Hepatitis C virus core protein epigenetically silences SFRP1 and enhances HCC aggressiveness by inducing epithelial-mesenchymal transition. | 2014 | Oncogene | pmid:23770846 |
| Wu Y et al. | Negative regulation of bcl-2 expression by p53 in hematopoietic cells. | 2001 | Oncogene | pmid:11313951 |
| Rashid SF et al. | Synergistic growth inhibition of prostate cancer cells by 1 alpha,25 Dihydroxyvitamin D(3) and its 19-nor-hexafluoride analogs in combination with either sodium butyrate or trichostatin A. | 2001 | Oncogene | pmid:11313934 |
| Lu Z et al. | E2F-HDAC complexes negatively regulate the tumor suppressor gene ARHI in breast cancer. | 2006 | Oncogene | pmid:16158053 |
| Hajji N et al. | Combinatorial action of the HDAC inhibitor trichostatin A and etoposide induces caspase-mediated AIF-dependent apoptotic cell death in non-small cell lung carcinoma cells. | 2008 | Oncogene | pmid:18071312 |
| Mühlisch J et al. | Epigenetic repression of RASSF1A but not CASP8 in supratentorial PNET (sPNET) and atypical teratoid/rhabdoid tumors (AT/RT) of childhood. | 2006 | Oncogene | pmid:16186793 |
| Cismasiu VB et al. | BCL11B functionally associates with the NuRD complex in T lymphocytes to repress targeted promoter. | 2005 | Oncogene | pmid:16091750 |
| Jiemjit A et al. | p21(WAF1/CIP1) induction by 5-azacytosine nucleosides requires DNA damage. | 2008 | Oncogene | pmid:18223691 |
| Cras A et al. | Epigenetic patterns of the retinoic acid receptor beta2 promoter in retinoic acid-resistant thyroid cancer cells. | 2007 | Oncogene | pmid:17213810 |
| Karagiannis TC et al. | Disparity of histone deacetylase inhibition on repair of radiation-induced DNA damage on euchromatin and constitutive heterochromatin compartments. | 2007 | Oncogene | pmid:17213813 |
| Sirchia SM et al. | Evidence of epigenetic changes affecting the chromatin state of the retinoic acid receptor beta2 promoter in breast cancer cells. | 2000 | Oncogene | pmid:10734315 |
| Toyooka S et al. | Progressive aberrant methylation of the RASSF1A gene in simian virus 40 infected human mesothelial cells. | 2002 | Oncogene | pmid:12082623 |
| Sengupta S et al. | Tumor suppressor p53 represses transcription of RECQ4 helicase. | 2005 | Oncogene | pmid:15674334 |
| Busson M et al. | Coactivation of nuclear receptors and myogenic factors induces the major BTG1 influence on muscle differentiation. | 2005 | Oncogene | pmid:15674337 |
| Jang ER et al. | The histone deacetylase inhibitor trichostatin A sensitizes estrogen receptor alpha-negative breast cancer cells to tamoxifen. | 2004 | Oncogene | pmid:14676837 |
| Kostyniuk CL et al. | The ubiquitous and tissue specific promoters of the human SRC gene are repressed by inhibitors of histone deacetylases. | 2002 | Oncogene | pmid:12214274 |
| López-Soto A et al. | HDAC3 represses the expression of NKG2D ligands ULBPs in epithelial tumour cells: potential implications for the immunosurveillance of cancer. | 2009 | Oncogene | pmid:19430493 |
| Reid G et al. | Multiple mechanisms induce transcriptional silencing of a subset of genes, including oestrogen receptor alpha, in response to deacetylase inhibition by valproic acid and trichostatin A. | 2005 | Oncogene | pmid:15870696 |
| Sengupta S et al. | Human AP endonuclease (APE1/Ref-1) and its acetylation regulate YB-1-p300 recruitment and RNA polymerase II loading in the drug-induced activation of multidrug resistance gene MDR1. | 2011 | Oncogene | pmid:20856196 |
| Shim JS et al. | Plakoglobin is a new target gene of histone deacetylase in human fibrosarcoma HT1080 cells. | 2004 | Oncogene | pmid:14661058 |
| Fleury L et al. | Eliminating epigenetic barriers induces transient hormone-regulated gene expression in estrogen receptor negative breast cancer cells. | 2008 | Oncogene | pmid:18317449 |
| Kim YB et al. | Oxamflatin is a novel antitumor compound that inhibits mammalian histone deacetylase. | 1999 | Oncogene | pmid:10229197 |
| Shin HJ et al. | Inhibition of histone deacetylase activity increases chromosomal instability by the aberrant regulation of mitotic checkpoint activation. | 2003 | Oncogene | pmid:12813458 |
| Kim TY et al. | Transcriptional silencing of the DLC-1 tumor suppressor gene by epigenetic mechanism in gastric cancer cells. | 2003 | Oncogene | pmid:12813468 |
| Suzuki M et al. | Direct association between PU.1 and MeCP2 that recruits mSin3A-HDAC complex for PU.1-mediated transcriptional repression. | 2003 | Oncogene | pmid:14647463 |
| Coombes MM et al. | Resetting the histone code at CDKN2A in HNSCC by inhibition of DNA methylation. | 2003 | Oncogene | pmid:14654786 |
| Chan AS et al. | Downregulation of ID4 by promoter hypermethylation in gastric adenocarcinoma. | 2003 | Oncogene | pmid:14534543 |
| Krishnan M et al. | HDAC inhibitors regulate claudin-1 expression in colon cancer cells through modulation of mRNA stability. | 2010 | Oncogene | pmid:19881542 |
| Shteper PJ et al. | Role of promoter methylation in regulation of the mammalian heparanase gene. | 2003 | Oncogene | pmid:14586400 |
| Hirose T et al. | p53-independent induction of Gadd45 by histone deacetylase inhibitor: coordinate regulation by transcription factors Oct-1 and NF-Y. | 2003 | Oncogene | pmid:14586402 |
| Vaghefi H and Neet KE | Deacetylation of p53 after nerve growth factor treatment in PC12 cells as a post-translational modification mechanism of neurotrophin-induced tumor suppressor activation. | 2004 | Oncogene | pmid:15361854 |
| Ibanez de Caceres I et al. | IGFBP-3 hypermethylation-derived deficiency mediates cisplatin resistance in non-small-cell lung cancer. | 2010 | Oncogene | pmid:20023704 |
| Zubia A et al. | Identification of (1H)-pyrroles as histone deacetylase inhibitors with antitumoral activity. | 2009 | Oncogene | pmid:19169274 |
| Lund P et al. | Oncogenic HRAS suppresses clusterin expression through promoter hypermethylation. | 2006 | Oncogene | pmid:16568090 |
| Sato N et al. | Epigenetic inactivation of TFPI-2 as a common mechanism associated with growth and invasion of pancreatic ductal adenocarcinoma. | 2005 | Oncogene | pmid:15592528 |