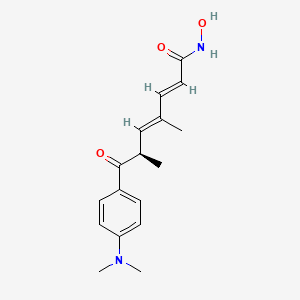
trichostatin A
Trichostatin is a lipid of Polyketides (PK) class. Trichostatin is associated with abnormalities such as Dentatorubral-Pallidoluysian Atrophy, PARAGANGLIOMAS 3, abnormal fragmented structure, Disintegration (morphologic abnormality) and Hyperostosis, Diffuse Idiopathic Skeletal. The involved functions are known as Acetylation, Cell Differentiation process, histone modification, Gene Silencing and Transcriptional Activation. Trichostatin often locates in CD41a, Hematopoietic System, Chromatin Structure, Blood and Endothelium. The associated genes with Trichostatin are SPI1 gene, CELL Gene, Chromatin, CXCR4 gene and DNMT1 gene. The related lipids are Butyrates, Promega, butyrate, Lipopolysaccharides and Steroids. The related experimental models are Knock-out, Mouse Model, Xenograft Model and Cancer Model.
Cross Reference
Introduction
To understand associated biological information of trichostatin A, we collected biological information of abnormalities, associated pathways, cellular/molecular locations, biological functions, related genes/proteins, lipids and common seen animal/experimental models with organized paragraphs from literatures.
What diseases are associated with trichostatin A?
trichostatin A is suspected in Infection, Morphologically altered structure, Ureteral obstruction, Photosensitization, Atherosclerosis, Hypertrophic Cardiomyopathy and other diseases in descending order of the highest number of associated sentences.
Related references are mostly published in these journals:
| Disease | Cross reference | Weighted score | Related literature |
|---|
Possible diseases from mapped MeSH terms on references
We collected disease MeSH terms mapped to the references associated with trichostatin A
PubChem Associated disorders and diseases
What pathways are associated with trichostatin A
Lipid pathways are not clear in current pathway databases. We organized associated pathways with trichostatin A through full-text articles, including metabolic pathways or pathways of biological mechanisms.
Related references are published most in these journals:
| Pathway name | Related literatures |
|---|
PubChem Biomolecular Interactions and Pathways
Link to PubChem Biomolecular Interactions and PathwaysWhat cellular locations are associated with trichostatin A?
Visualization in cellular structure
Associated locations are in red color. Not associated locations are in black.
Related references are published most in these journals:
| Location | Cross reference | Weighted score | Related literatures |
|---|
What functions are associated with trichostatin A?
Related references are published most in these journals:
| Function | Cross reference | Weighted score | Related literatures |
|---|
What lipids are associated with trichostatin A?
Related references are published most in these journals:
| Lipid concept | Cross reference | Weighted score | Related literatures |
|---|
What genes are associated with trichostatin A?
Related references are published most in these journals:
| Gene | Cross reference | Weighted score | Related literatures |
|---|
What common seen animal models are associated with trichostatin A?
Mouse Model
Mouse Model are used in the study 'Regulation of minichromosome maintenance gene family by microRNA-1296 and genistein in prostate cancer.' (Majid S et al., 2010), Mouse Model are used in the study 'Reversal of hypermethylation and reactivation of p16INK4a, RARbeta, and MGMT genes by genistein and other isoflavones from soy.' (Fang MZ et al., 2005) and Mouse Model are used in the study 'Histone deacetylase 3 mediates allergic skin inflammation by regulating expression of MCP1 protein.' (Kim Y et al., 2012).
Xenograft Model
Xenograft Model are used in the study 'Histone deacetylase inhibitors induce growth arrest and differentiation in uveal melanoma.' (Landreville S et al., 2012), Xenograft Model are used in the study 'Extended treatment with physiologic concentrations of dietary phytochemicals results in altered gene expression, reduced growth, and apoptosis of cancer cells.' (Moiseeva EP et al., 2007) and Xenograft Model are used in the study 'Retinoic acid and the histone deacetylase inhibitor trichostatin a inhibit the proliferation of human renal cell carcinoma in a xenograft tumor model.' (Touma SE et al., 2005).
Cancer Model
Cancer Model are used in the study 'Plasma pharmacokinetics and metabolism of the histone deacetylase inhibitor trichostatin a after intraperitoneal administration to mice.' (Sanderson L et al., 2004).
Related references are published most in these journals:
| Model | Cross reference | Weighted score | Related literatures |
|---|
NCBI Entrez Crosslinks
All references with trichostatin A
Download all related citations| Authors | Title | Published | Journal | PubMed Link |
|---|---|---|---|---|
| Henderson C et al. | Role of caspases, Bid, and p53 in the apoptotic response triggered by histone deacetylase inhibitors trichostatin-A (TSA) and suberoylanilide hydroxamic acid (SAHA). | 2003 | J. Biol. Chem. | pmid:12556448 |
| Huang W et al. | Trichostatin A induces transforming growth factor beta type II receptor promoter activity and acetylation of Sp1 by recruitment of PCAF/p300 to a Sp1.NF-Y complex. | 2005 | J. Biol. Chem. | pmid:15647279 |
| Galasinski SC et al. | Global regulation of post-translational modifications on core histones. | 2002 | J. Biol. Chem. | pmid:11709551 |
| Corvetta D et al. | Physical interaction between MYCN oncogene and polycomb repressive complex 2 (PRC2) in neuroblastoma: functional and therapeutic implications. | 2013 | J. Biol. Chem. | pmid:23362253 |
| Yin L et al. | Butyrate suppression of colonocyte NF-kappa B activation and cellular proteasome activity. | 2001 | J. Biol. Chem. | pmid:11572859 |
| Stimson KM and Vertino PM | Methylation-mediated silencing of TMS1/ASC is accompanied by histone hypoacetylation and CpG island-localized changes in chromatin architecture. | 2002 | J. Biol. Chem. | pmid:11733524 |
| Sakai N et al. | Involvement of histone acetylation in ovarian steroid-induced decidualization of human endometrial stromal cells. | 2003 | J. Biol. Chem. | pmid:12609987 |
| Wang S and Zhu J | Evidence for a relief of repression mechanism for activation of the human telomerase reverse transcriptase promoter. | 2003 | J. Biol. Chem. | pmid:12611896 |
| Kurtev V et al. | Transcriptional regulation by the repressor of estrogen receptor activity via recruitment of histone deacetylases. | 2004 | J. Biol. Chem. | pmid:15140878 |
| Klampfer L et al. | Requirement of histone deacetylase activity for signaling by STAT1. | 2004 | J. Biol. Chem. | pmid:15123634 |
| Fu M et al. | The androgen receptor acetylation site regulates cAMP and AKT but not ERK-induced activity. | 2004 | J. Biol. Chem. | pmid:15123687 |
| Rigamonti D et al. | Loss of huntingtin function complemented by small molecules acting as repressor element 1/neuron restrictive silencer element silencer modulators. | 2007 | J. Biol. Chem. | pmid:17565993 |
| Roy S and Tenniswood M | Site-specific acetylation of p53 directs selective transcription complex assembly. | 2007 | J. Biol. Chem. | pmid:17121856 |
| Jin YH et al. | Transforming growth factor-beta stimulates p300-dependent RUNX3 acetylation, which inhibits ubiquitination-mediated degradation. | 2004 | J. Biol. Chem. | pmid:15138260 |
| Milutinovic S et al. | DNA methyltransferase 1 knock down induces gene expression by a mechanism independent of DNA methylation and histone deacetylation. | 2004 | J. Biol. Chem. | pmid:15087453 |
| Butler PL et al. | Acetylation stimulates the epithelial sodium channel by reducing its ubiquitination and degradation. | 2015 | J. Biol. Chem. | pmid:25787079 |
| Gusterson RJ et al. | The transcriptional co-activators CREB-binding protein (CBP) and p300 play a critical role in cardiac hypertrophy that is dependent on their histone acetyltransferase activity. | 2003 | J. Biol. Chem. | pmid:12477714 |
| Phiel CJ et al. | Histone deacetylase is a direct target of valproic acid, a potent anticonvulsant, mood stabilizer, and teratogen. | 2001 | J. Biol. Chem. | pmid:11473107 |
| Jacob AL et al. | Acetylation of steroidogenic factor 1 protein regulates its transcriptional activity and recruits the coactivator GCN5. | 2001 | J. Biol. Chem. | pmid:11479297 |
| Gregory RI et al. | Inhibition of histone deacetylases alters allelic chromatin conformation at the imprinted U2af1-rs1 locus in mouse embryonic stem cells. | 2002 | J. Biol. Chem. | pmid:11821379 |
| Lim Y et al. | Trichostatin A-induced detransformation correlates with decreased focal adhesion kinase phosphorylation at tyrosine 861 in ras-transformed fibroblasts. | 2002 | J. Biol. Chem. | pmid:11821402 |
| Wilson MA et al. | The histone deacetylase inhibitor trichostatin A blocks progesterone receptor-mediated transactivation of the mouse mammary tumor virus promoter in vivo. | 2002 | J. Biol. Chem. | pmid:11821430 |
| Brush MH et al. | Deactylase inhibitors disrupt cellular complexes containing protein phosphatases and deacetylases. | 2004 | J. Biol. Chem. | pmid:14670976 |
| Qiao Y et al. | Dual roles of histone H3 lysine 9 acetylation in human embryonic stem cell pluripotency and neural differentiation. | 2015 | J. Biol. Chem. | pmid:25519907 |
| Hattori N et al. | Epigenetic control of mouse Oct-4 gene expression in embryonic stem cells and trophoblast stem cells. | 2004 | J. Biol. Chem. | pmid:14761969 |
| Horion J et al. | Histone deacetylase inhibitor trichostatin A sustains sodium pervanadate-induced NF-kappaB activation by delaying IkappaBalpha mRNA resynthesis: comparison with tumor necrosis factor alpha. | 2007 | J. Biol. Chem. | pmid:17409387 |
| Christian M et al. | Characterization of four autonomous repression domains in the corepressor receptor interacting protein 140. | 2004 | J. Biol. Chem. | pmid:14736873 |
| Bu Y and Gelman IH | v-Src-mediated down-regulation of SSeCKS metastasis suppressor gene promoter by the recruitment of HDAC1 into a USF1-Sp1-Sp3 complex. | 2007 | J. Biol. Chem. | pmid:17626016 |
| Park SH et al. | Transcriptional regulation of the transforming growth factor beta type II receptor gene by histone acetyltransferase and deacetylase is mediated by NF-Y in human breast cancer cells. | 2002 | J. Biol. Chem. | pmid:11744689 |
| Wang Y et al. | Nuclear factor κB mediates suppression of canonical transient receptor potential 6 expression by reactive oxygen species and protein kinase C in kidney cells. | 2013 | J. Biol. Chem. | pmid:23525112 |
| Singal R et al. | Methylation of promoter proximal-transcribed sequences of an embryonic globin gene inhibits transcription in primary erythroid cells and promotes formation of a cell type-specific methyl cytosine binding complex. | 2002 | J. Biol. Chem. | pmid:11684679 |
| Ammanamanchi S et al. | Acetylated sp3 is a transcriptional activator. | 2003 | J. Biol. Chem. | pmid:12837748 |
| Seth KA and Majzoub JA | Repressor element silencing transcription factor/neuron-restrictive silencing factor (REST/NRSF) can act as an enhancer as well as a repressor of corticotropin-releasing hormone gene transcription. | 2001 | J. Biol. Chem. | pmid:11278361 |
| Xiao H et al. | p300 collaborates with Sp1 and Sp3 in p21(waf1/cip1) promoter activation induced by histone deacetylase inhibitor. | 2000 | J. Biol. Chem. | pmid:10625687 |
| Cribbs A et al. | Inhibition of histone H3K27 demethylases selectively modulates inflammatory phenotypes of natural killer cells. | 2018 | J. Biol. Chem. | pmid:29301935 |
| Wang J et al. | Histone deacetylase inhibitors suppress TF-kappaB-dependent agonist-driven tissue factor expression in endothelial cells and monocytes. | 2007 | J. Biol. Chem. | pmid:17675290 |
| Sarg B et al. | Histone H4 hyperacetylation precludes histone H4 lysine 20 trimethylation. | 2004 | J. Biol. Chem. | pmid:15456746 |
| Yoshida T et al. | Kruppel-like factor 4 protein regulates isoproterenol-induced cardiac hypertrophy by modulating myocardin expression and activity. | 2014 | J. Biol. Chem. | pmid:25100730 |
| Hishiki T et al. | BCL3 acts as a negative regulator of transcription from the human T-cell leukemia virus type 1 long terminal repeat through interactions with TORC3. | 2007 | J. Biol. Chem. | pmid:17644518 |
| Yoshida M et al. | Potent and specific inhibition of mammalian histone deacetylase both in vivo and in vitro by trichostatin A. | 1990 | J. Biol. Chem. | pmid:2211619 |
| Lin AC et al. | The N termini of Friend of GATA (FOG) proteins define a novel transcriptional repression motif and a superfamily of transcriptional repressors. | 2004 | J. Biol. Chem. | pmid:15507435 |
| Kekatpure VD et al. | HDAC6 modulates Hsp90 chaperone activity and regulates activation of aryl hydrocarbon receptor signaling. | 2009 | J. Biol. Chem. | pmid:19158084 |
| Chang S and Pikaard CS | Transcript profiling in Arabidopsis reveals complex responses to global inhibition of DNA methylation and histone deacetylation. | 2005 | J. Biol. Chem. | pmid:15516340 |
| Wang S and Zhu J | The hTERT gene is embedded in a nuclease-resistant chromatin domain. | 2004 | J. Biol. Chem. | pmid:15516693 |
| Lauffer BE et al. | Histone deacetylase (HDAC) inhibitor kinetic rate constants correlate with cellular histone acetylation but not transcription and cell viability. | 2013 | J. Biol. Chem. | pmid:23897821 |
| Mogal AP et al. | Haploinsufficient prostate tumor suppression by Nkx3.1: a role for chromatin accessibility in dosage-sensitive gene regulation. | 2007 | J. Biol. Chem. | pmid:17602165 |
| Wong CY et al. | Epigenetic regulation of phosphatidylinositol 3,4,5-triphosphate-dependent Rac exchanger 1 gene expression in prostate cancer cells. | 2011 | J. Biol. Chem. | pmid:21636851 |
| Asoh S et al. | The super anti-apoptotic factor Bcl-xFNK constructed by disturbing intramolecular polar interactions in rat Bcl-xL. | 2000 | J. Biol. Chem. | pmid:10970895 |
| Cong YS and Bacchetti S | Histone deacetylation is involved in the transcriptional repression of hTERT in normal human cells. | 2000 | J. Biol. Chem. | pmid:10986277 |
| Wei LN et al. | Receptor-interacting protein 140 directly recruits histone deacetylases for gene silencing. | 2000 | J. Biol. Chem. | pmid:11006275 |
| Chen C et al. | Evidence that silencing of the HPRT promoter by DNA methylation is mediated by critical CpG sites. | 2001 | J. Biol. Chem. | pmid:11013250 |
| Su M et al. | Recruitment of nuclear factor Y to the inverted CCAAT element (ICE) by c-Jun and E1A stimulates basal transcription of the bone sialoprotein gene in osteosarcoma cells. | 2005 | J. Biol. Chem. | pmid:16087680 |
| Kadiyala V et al. | Class I lysine deacetylases facilitate glucocorticoid-induced transcription. | 2013 | J. Biol. Chem. | pmid:23946490 |
| Milutinovic S et al. | Proliferating cell nuclear antigen associates with histone deacetylase activity, integrating DNA replication and chromatin modification. | 2002 | J. Biol. Chem. | pmid:11929879 |
| McCool KW et al. | The role of histone acetylation in regulating early gene expression patterns during early embryonic stem cell differentiation. | 2007 | J. Biol. Chem. | pmid:17204470 |
| Wen J et al. | Tal1/SCL binding to pericentromeric DNA represses transcription. | 2005 | J. Biol. Chem. | pmid:15677454 |
| Kim Y et al. | Histone deacetylase 3 mediates allergic skin inflammation by regulating expression of MCP1 protein. | 2012 | J. Biol. Chem. | pmid:22679019 |
| Waterborg JH | Dynamics of histone acetylation in Chlamydomonas reinhardtii. | 1998 | J. Biol. Chem. | pmid:9765294 |
| Kizuka Y et al. | Epigenetic regulation of a brain-specific glycosyltransferase N-acetylglucosaminyltransferase-IX (GnT-IX) by specific chromatin modifiers. | 2014 | J. Biol. Chem. | pmid:24619417 |
| Chang I and Wang CY | Inhibition of HDAC6 Protein Enhances Bortezomib-induced Apoptosis in Head and Neck Squamous Cell Carcinoma (HNSCC) by Reducing Autophagy. | 2016 | J. Biol. Chem. | pmid:27369083 |
| Verdel A and Khochbin S | Identification of a new family of higher eukaryotic histone deacetylases. Coordinate expression of differentiation-dependent chromatin modifiers. | 1999 | J. Biol. Chem. | pmid:9891014 |
| Simboeck E et al. | A phosphorylation switch regulates the transcriptional activation of cell cycle regulator p21 by histone deacetylase inhibitors. | 2010 | J. Biol. Chem. | pmid:20952396 |
| Yamaguchi K et al. | Histone deacetylase inhibitors suppress the induction of c-Jun and its target genes including COX-2. | 2005 | J. Biol. Chem. | pmid:15994313 |
| Wooten LG and Ogretmen B | Sp1/Sp3-dependent regulation of human telomerase reverse transcriptase promoter activity by the bioactive sphingolipid ceramide. | 2005 | J. Biol. Chem. | pmid:15951564 |
| Marks J et al. | Two functionally divergent p53-responsive elements in the rat bradykinin B2 receptor promoter. | 2003 | J. Biol. Chem. | pmid:12791684 |
| Vanden Berghe W et al. | The nuclear factor-kappaB engages CBP/p300 and histone acetyltransferase activity for transcriptional activation of the interleukin-6 gene promoter. | 1999 | J. Biol. Chem. | pmid:10542243 |
| Murumägi A et al. | Characterization of regulatory elements and methylation pattern of the autoimmune regulator (AIRE) promoter. | 2003 | J. Biol. Chem. | pmid:12651856 |
| Liao M et al. | Coactivator function of positive cofactor 4 (PC4) in Sp1-directed luteinizing hormone receptor (LHR) gene transcription. | 2011 | J. Biol. Chem. | pmid:21193408 |
| Liu CJ et al. | Leukemia/lymphoma-related factor, a POZ domain-containing transcriptional repressor, interacts with histone deacetylase-1 and inhibits cartilage oligomeric matrix protein gene expression and chondrogenesis. | 2004 | J. Biol. Chem. | pmid:15337766 |
| Muralidhar SA et al. | Histone deacetylase 9 activates gamma-globin gene expression in primary erythroid cells. | 2011 | J. Biol. Chem. | pmid:21078662 |
| Hayakawa F et al. | Acetylation of PML is involved in histone deacetylase inhibitor-mediated apoptosis. | 2008 | J. Biol. Chem. | pmid:18621739 |
| Kawamura T et al. | Acetylation of GATA-4 is involved in the differentiation of embryonic stem cells into cardiac myocytes. | 2005 | J. Biol. Chem. | pmid:15764815 |
| Webster SJ et al. | Regulation of GTP-binding protein (Gαs) expression in human myometrial cells: a role for tumor necrosis factor in modulating Gαs promoter acetylation by transcriptional complexes. | 2013 | J. Biol. Chem. | pmid:23297421 |
| Johnson CA et al. | Deacetylase activity associates with topoisomerase II and is necessary for etoposide-induced apoptosis. | 2001 | J. Biol. Chem. | pmid:11136718 |
| Vos MD et al. | The pro-apoptotic Ras effector Nore1 may serve as a Ras-regulated tumor suppressor in the lung. | 2003 | J. Biol. Chem. | pmid:12676952 |
| Yoshioka H et al. | Nonsteroidal anti-inflammatory drug-activated gene (NAG-1/GDF15) expression is increased by the histone deacetylase inhibitor trichostatin A. | 2008 | J. Biol. Chem. | pmid:18801729 |
| Banchio C et al. | Role of histone deacetylase in the expression of CTP:phosphocholine cytidylyltransferase alpha. | 2006 | J. Biol. Chem. | pmid:16484221 |
| Sakamoto S et al. | Histone deacetylase activity is required to recruit RNA polymerase II to the promoters of selected interferon-stimulated early response genes. | 2004 | J. Biol. Chem. | pmid:15194680 |
| Ng Y et al. | Expression of the human myotonic dystrophy kinase-related Cdc42-binding kinase gamma is regulated by promoter DNA methylation and Sp1 binding. | 2004 | J. Biol. Chem. | pmid:15194684 |
| Theisen JW et al. | Biochemical analysis of histone deacetylase-independent transcriptional repression by MeCP2. | 2013 | J. Biol. Chem. | pmid:23349465 |
| Zhang Y and Zhang B | Trichostatin A, an Inhibitor of Histone Deacetylase, Inhibits the Viability and Invasiveness of Hypoxic Rheumatoid Arthritis Fibroblast-Like Synoviocytes via PI3K/Akt Signaling. | 2016 | J. Biochem. Mol. Toxicol. | pmid:26509796 |
| Jin B and Ryu DY | Regulation of CYP1A2 by histone deacetylase inhibitors in mouse hepatocytes. | 2004 | J. Biochem. Mol. Toxicol. | pmid:15252868 |
| Kook SH et al. | Epstein-Barr virus-infected Akata cells are sensitive to histone deacetylase inhibitor TSA-provoked apoptosis. | 2005 | J. Biochem. Mol. Biol. | pmid:16336792 |
| Maruoka H et al. | Low-molecular-weight compounds having neurotrophic activity in cultured PC12 cells and neurons. | 2011 | J. Biochem. | pmid:21908547 |
| Yamamoto I et al. | Histone hyperacetylation plays a role in augmentation of IL-4-induced IgE production in LPS-stimulated murine B-lymphocytes by sodium butyrate. | 1996 | J. Biochem. | pmid:8827437 |
| Zhao W et al. | The essential role of histone H3 Lys9 di-methylation and MeCP2 binding in MGMT silencing with poor DNA methylation of the promoter CpG island. | 2005 | J. Biochem. | pmid:15809347 |
| Hildmann C et al. | A new amidohydrolase from Bordetella or Alcaligenes strain FB188 with similarities to histone deacetylases. | 2004 | J. Bacteriol. | pmid:15060035 |
| Reilly CM et al. | The histone deacetylase inhibitor trichostatin A upregulates regulatory T cells and modulates autoimmunity in NZB/W F1 mice. | 2008 | J. Autoimmun. | pmid:18650065 |
| Azechi T et al. | Trichostatin A, an HDAC class I/II inhibitor, promotes Pi-induced vascular calcification via up-regulation of the expression of alkaline phosphatase. | 2013 | J. Atheroscler. Thromb. | pmid:23518467 |
| Okamoto H et al. | Trichostatin A, an inhibitor of histone deacetylase, inhibits smooth muscle cell proliferation via induction of p21(WAF1). | 2006 | J. Atheroscler. Thromb. | pmid:16908950 |
| Slattery EL et al. | Epigenetic influences on sensory regeneration: histone deacetylases regulate supporting cell proliferation in the avian utricle. | 2009 | J. Assoc. Res. Otolaryngol. | pmid:19340485 |
| Jafari S et al. | Epigenetic modification does not determine the time of POU5F1 transcription activation in cloned bovine embryos. | 2011 | J. Assist. Reprod. Genet. | pmid:22020531 |
| Dupré-Aucouturier S et al. | Trichostatin A, a histone deacetylase inhibitor, modulates unloaded-induced skeletal muscle atrophy. | 2015 | J. Appl. Physiol. | pmid:26112243 |
| Ueda JY et al. | JBIR-17, a novel trichostatin analog from Streptomyces sp. 26634. | 2009 | J. Antibiot. | pmid:19300468 |
| Yoshida M et al. | Structural specificity for biological activity of trichostatin A, a specific inhibitor of mammalian cell cycle with potent differentiation-inducing activity in Friend leukemia cells. | 1990 | J. Antibiot. | pmid:2211374 |
| Otoguro K et al. | Screening for new antitrichomonal substances of microbial origin and antitrichomonal activity of trichostatin A. | 1988 | J. Antibiot. | pmid:3372352 |
| Takahashi I et al. | Selective inhibition of IL-2 gene expression by trichostatin A, a potent inhibitor of mammalian histone deacetylase. | 1996 | J. Antibiot. | pmid:8682722 |
| Komatsu Y and Hayashi H | Histone deacetylase inhibitors up-regulate the expression of cell surface MHC class-I molecules in B16/BL6 cells. | 1998 | J. Antibiot. | pmid:9531994 |
| Koguchi Y et al. | Trichostatin A and herboxidiene up-regulate the gene expression of low density lipoprotein receptor. | 1997 | J. Antibiot. | pmid:9592573 |
| Kim YB et al. | Mechanism of cell cycle arrest caused by histone deacetylase inhibitors in human carcinoma cells. | 2000 | J. Antibiot. | pmid:11132966 |