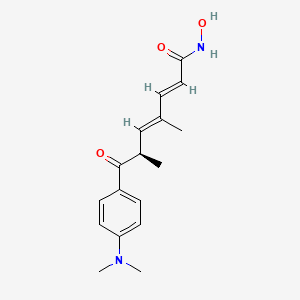
trichostatin A
Trichostatin is a lipid of Polyketides (PK) class. Trichostatin is associated with abnormalities such as Dentatorubral-Pallidoluysian Atrophy, PARAGANGLIOMAS 3, abnormal fragmented structure, Disintegration (morphologic abnormality) and Hyperostosis, Diffuse Idiopathic Skeletal. The involved functions are known as Acetylation, Cell Differentiation process, histone modification, Gene Silencing and Transcriptional Activation. Trichostatin often locates in CD41a, Hematopoietic System, Chromatin Structure, Blood and Endothelium. The associated genes with Trichostatin are SPI1 gene, CELL Gene, Chromatin, CXCR4 gene and DNMT1 gene. The related lipids are Butyrates, Promega, butyrate, Lipopolysaccharides and Steroids. The related experimental models are Knock-out, Mouse Model, Xenograft Model and Cancer Model.
Cross Reference
Introduction
To understand associated biological information of trichostatin A, we collected biological information of abnormalities, associated pathways, cellular/molecular locations, biological functions, related genes/proteins, lipids and common seen animal/experimental models with organized paragraphs from literatures.
What diseases are associated with trichostatin A?
trichostatin A is suspected in Infection, Morphologically altered structure, Ureteral obstruction, Photosensitization, Atherosclerosis, Hypertrophic Cardiomyopathy and other diseases in descending order of the highest number of associated sentences.
Related references are mostly published in these journals:
| Disease | Cross reference | Weighted score | Related literature |
|---|
Possible diseases from mapped MeSH terms on references
We collected disease MeSH terms mapped to the references associated with trichostatin A
PubChem Associated disorders and diseases
What pathways are associated with trichostatin A
Lipid pathways are not clear in current pathway databases. We organized associated pathways with trichostatin A through full-text articles, including metabolic pathways or pathways of biological mechanisms.
Related references are published most in these journals:
| Pathway name | Related literatures |
|---|
PubChem Biomolecular Interactions and Pathways
Link to PubChem Biomolecular Interactions and PathwaysWhat cellular locations are associated with trichostatin A?
Visualization in cellular structure
Associated locations are in red color. Not associated locations are in black.
Related references are published most in these journals:
| Location | Cross reference | Weighted score | Related literatures |
|---|
What functions are associated with trichostatin A?
Related references are published most in these journals:
| Function | Cross reference | Weighted score | Related literatures |
|---|
What lipids are associated with trichostatin A?
Related references are published most in these journals:
| Lipid concept | Cross reference | Weighted score | Related literatures |
|---|
What genes are associated with trichostatin A?
Related references are published most in these journals:
| Gene | Cross reference | Weighted score | Related literatures |
|---|
What common seen animal models are associated with trichostatin A?
Mouse Model
Mouse Model are used in the study 'Regulation of minichromosome maintenance gene family by microRNA-1296 and genistein in prostate cancer.' (Majid S et al., 2010), Mouse Model are used in the study 'Reversal of hypermethylation and reactivation of p16INK4a, RARbeta, and MGMT genes by genistein and other isoflavones from soy.' (Fang MZ et al., 2005) and Mouse Model are used in the study 'Histone deacetylase 3 mediates allergic skin inflammation by regulating expression of MCP1 protein.' (Kim Y et al., 2012).
Xenograft Model
Xenograft Model are used in the study 'Histone deacetylase inhibitors induce growth arrest and differentiation in uveal melanoma.' (Landreville S et al., 2012), Xenograft Model are used in the study 'Extended treatment with physiologic concentrations of dietary phytochemicals results in altered gene expression, reduced growth, and apoptosis of cancer cells.' (Moiseeva EP et al., 2007) and Xenograft Model are used in the study 'Retinoic acid and the histone deacetylase inhibitor trichostatin a inhibit the proliferation of human renal cell carcinoma in a xenograft tumor model.' (Touma SE et al., 2005).
Cancer Model
Cancer Model are used in the study 'Plasma pharmacokinetics and metabolism of the histone deacetylase inhibitor trichostatin a after intraperitoneal administration to mice.' (Sanderson L et al., 2004).
Related references are published most in these journals:
| Model | Cross reference | Weighted score | Related literatures |
|---|
NCBI Entrez Crosslinks
All references with trichostatin A
Download all related citations| Authors | Title | Published | Journal | PubMed Link |
|---|---|---|---|---|
| Avram D et al. | Isolation of a novel family of C(2)H(2) zinc finger proteins implicated in transcriptional repression mediated by chicken ovalbumin upstream promoter transcription factor (COUP-TF) orphan nuclear receptors. | 2000 | J. Biol. Chem. | pmid:10744719 |
| Eickhoff B et al. | Trichostatin A modulates expression of p21waf1/cip1, Bcl-xL, ID1, ID2, ID3, CRAB2, GATA-2, hsp86 and TFIID/TAFII31 mRNA in human lung adenocarcinoma cells. | 2000 | Biol. Chem. | pmid:10746741 |
| Hobbs CA and Gilmour SK | High levels of intracellular polyamines promote histone acetyltransferase activity resulting in chromatin hyperacetylation. | 2000 | J. Cell. Biochem. | pmid:10760944 |
| Yoshida M and Horinouchi S | [Histone deacetylase inhibitors--new anticancer agents?]. | 2000 | Tanpakushitsu Kakusan Koso | pmid:10771678 |
| Yu F et al. | Histone deacetylase-independent transcriptional repression by methyl-CpG-binding protein 2. | 2000 | Nucleic Acids Res. | pmid:10773092 |
| Lee SK et al. | Silencing mediator of retinoic acid and thyroid hormone receptors, as a novel transcriptional corepressor molecule of activating protein-1, nuclear factor-kappaB, and serum response factor. | 2000 | J. Biol. Chem. | pmid:10777532 |
| Zhang HS et al. | Exit from G1 and S phase of the cell cycle is regulated by repressor complexes containing HDAC-Rb-hSWI/SNF and Rb-hSWI/SNF. | 2000 | Cell | pmid:10778858 |
| Fu M et al. | p300 and p300/cAMP-response element-binding protein-associated factor acetylate the androgen receptor at sites governing hormone-dependent transactivation. | 2000 | J. Biol. Chem. | pmid:10779504 |
| Kitamura K et al. | Histone deacetylase inhibitor but not arsenic trioxide differentiates acute promyelocytic leukaemia cells with t(11;17) in combination with all-trans retinoic acid. | 2000 | Br. J. Haematol. | pmid:10792271 |
| Gays F et al. | The mouse tumor cell lines EL4 and RMA display mosaic expression of NK-related and certain other surface molecules and appear to have a common origin. | 2000 | J. Immunol. | pmid:10799866 |