| MeSH term | MeSH ID | Detail |
|---|---|---|
| Biliary Tract Neoplasms | D001661 | 7 associated lipids |
| Autoimmune Diseases | D001327 | 27 associated lipids |
| Asthma | D001249 | 52 associated lipids |
| Arthritis, Experimental | D001169 | 24 associated lipids |
| Alzheimer Disease | D000544 | 76 associated lipids |
| Adrenoleukodystrophy | D000326 | 29 associated lipids |
| Adenocarcinoma, Papillary | D000231 | 2 associated lipids |
| Adenocarcinoma | D000230 | 166 associated lipids |
| Abnormalities, Multiple | D000015 | 13 associated lipids |
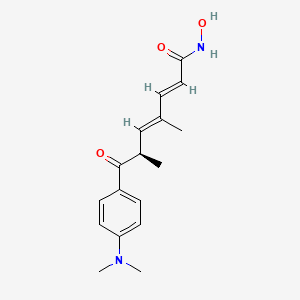
trichostatin A
Trichostatin is a lipid of Polyketides (PK) class. Trichostatin is associated with abnormalities such as Dentatorubral-Pallidoluysian Atrophy, PARAGANGLIOMAS 3, abnormal fragmented structure, Disintegration (morphologic abnormality) and Hyperostosis, Diffuse Idiopathic Skeletal. The involved functions are known as Acetylation, Cell Differentiation process, histone modification, Gene Silencing and Transcriptional Activation. Trichostatin often locates in CD41a, Hematopoietic System, Chromatin Structure, Blood and Endothelium. The associated genes with Trichostatin are SPI1 gene, CELL Gene, Chromatin, CXCR4 gene and DNMT1 gene. The related lipids are Butyrates, Promega, butyrate, Lipopolysaccharides and Steroids. The related experimental models are Knock-out, Mouse Model, Xenograft Model and Cancer Model.
Cross Reference
Introduction
To understand associated biological information of trichostatin A, we collected biological information of abnormalities, associated pathways, cellular/molecular locations, biological functions, related genes/proteins, lipids and common seen animal/experimental models with organized paragraphs from literatures.
What diseases are associated with trichostatin A?
trichostatin A is suspected in Infection, Morphologically altered structure, Ureteral obstruction, Photosensitization, Atherosclerosis, Hypertrophic Cardiomyopathy and other diseases in descending order of the highest number of associated sentences.
Related references are mostly published in these journals:
| Disease | Cross reference | Weighted score | Related literature |
|---|
Possible diseases from mapped MeSH terms on references
We collected disease MeSH terms mapped to the references associated with trichostatin A
PubChem Associated disorders and diseases
What pathways are associated with trichostatin A
Lipid pathways are not clear in current pathway databases. We organized associated pathways with trichostatin A through full-text articles, including metabolic pathways or pathways of biological mechanisms.
Related references are published most in these journals:
| Pathway name | Related literatures |
|---|
PubChem Biomolecular Interactions and Pathways
Link to PubChem Biomolecular Interactions and PathwaysWhat cellular locations are associated with trichostatin A?
Visualization in cellular structure
Associated locations are in red color. Not associated locations are in black.
Related references are published most in these journals:
| Location | Cross reference | Weighted score | Related literatures |
|---|
What functions are associated with trichostatin A?
Related references are published most in these journals:
| Function | Cross reference | Weighted score | Related literatures |
|---|
What lipids are associated with trichostatin A?
Related references are published most in these journals:
| Lipid concept | Cross reference | Weighted score | Related literatures |
|---|
What genes are associated with trichostatin A?
Related references are published most in these journals:
| Gene | Cross reference | Weighted score | Related literatures |
|---|
What common seen animal models are associated with trichostatin A?
Mouse Model
Mouse Model are used in the study 'Regulation of minichromosome maintenance gene family by microRNA-1296 and genistein in prostate cancer.' (Majid S et al., 2010), Mouse Model are used in the study 'Reversal of hypermethylation and reactivation of p16INK4a, RARbeta, and MGMT genes by genistein and other isoflavones from soy.' (Fang MZ et al., 2005) and Mouse Model are used in the study 'Histone deacetylase 3 mediates allergic skin inflammation by regulating expression of MCP1 protein.' (Kim Y et al., 2012).
Xenograft Model
Xenograft Model are used in the study 'Histone deacetylase inhibitors induce growth arrest and differentiation in uveal melanoma.' (Landreville S et al., 2012), Xenograft Model are used in the study 'Extended treatment with physiologic concentrations of dietary phytochemicals results in altered gene expression, reduced growth, and apoptosis of cancer cells.' (Moiseeva EP et al., 2007) and Xenograft Model are used in the study 'Retinoic acid and the histone deacetylase inhibitor trichostatin a inhibit the proliferation of human renal cell carcinoma in a xenograft tumor model.' (Touma SE et al., 2005).
Cancer Model
Cancer Model are used in the study 'Plasma pharmacokinetics and metabolism of the histone deacetylase inhibitor trichostatin a after intraperitoneal administration to mice.' (Sanderson L et al., 2004).
Related references are published most in these journals:
| Model | Cross reference | Weighted score | Related literatures |
|---|
NCBI Entrez Crosslinks
All references with trichostatin A
Download all related citations| Authors | Title | Published | Journal | PubMed Link |
|---|---|---|---|---|
| Dar RD et al. | Transcriptional burst frequency and burst size are equally modulated across the human genome. | 2012 | Proc. Natl. Acad. Sci. U.S.A. | pmid:23064634 |
| Grisolano JL et al. | An activated receptor tyrosine kinase, TEL/PDGFbetaR, cooperates with AML1/ETO to induce acute myeloid leukemia in mice. | 2003 | Proc. Natl. Acad. Sci. U.S.A. | pmid:12881486 |
| Xu D et al. | Switch from Myc/Max to Mad1/Max binding and decrease in histone acetylation at the telomerase reverse transcriptase promoter during differentiation of HL60 cells. | 2001 | Proc. Natl. Acad. Sci. U.S.A. | pmid:11274400 |
| Magdinier F and Wolffe AP | Selective association of the methyl-CpG binding protein MBD2 with the silent p14/p16 locus in human neoplasia. | 2001 | Proc. Natl. Acad. Sci. U.S.A. | pmid:11309512 |
| Mishra N et al. | Trichostatin A reverses skewed expression of CD154, interleukin-10, and interferon-gamma gene and protein expression in lupus T cells. | 2001 | Proc. Natl. Acad. Sci. U.S.A. | pmid:11226290 |
| Zhang L et al. | Small molecule regulators of autophagy identified by an image-based high-throughput screen. | 2007 | Proc. Natl. Acad. Sci. U.S.A. | pmid:18024584 |
| Minucci S et al. | A histone deacetylase inhibitor potentiates retinoid receptor action in embryonal carcinoma cells. | 1997 | Proc. Natl. Acad. Sci. U.S.A. | pmid:9326603 |
| Duffy S et al. | Overexpression screens identify conserved dosage chromosome instability genes in yeast and human cancer. | 2016 | Proc. Natl. Acad. Sci. U.S.A. | pmid:27551064 |
| Iezzi S et al. | Stage-specific modulation of skeletal myogenesis by inhibitors of nuclear deacetylases. | 2002 | Proc. Natl. Acad. Sci. U.S.A. | pmid:12032356 |
| Ito K et al. | A molecular mechanism of action of theophylline: Induction of histone deacetylase activity to decrease inflammatory gene expression. | 2002 | Proc. Natl. Acad. Sci. U.S.A. | pmid:12070353 |
| Matoba S et al. | RNAi-mediated knockdown of Xist can rescue the impaired postimplantation development of cloned mouse embryos. | 2011 | Proc. Natl. Acad. Sci. U.S.A. | pmid:22065773 |
| Crump NT et al. | Dynamic acetylation of all lysine-4 trimethylated histone H3 is evolutionarily conserved and mediated by p300/CBP. | 2011 | Proc. Natl. Acad. Sci. U.S.A. | pmid:21518915 |
| Yeo M et al. | Bisphenol A delays the perinatal chloride shift in cortical neurons by epigenetic effects on the Kcc2 promoter. | 2013 | Proc. Natl. Acad. Sci. U.S.A. | pmid:23440186 |
| Saito A et al. | A synthetic inhibitor of histone deacetylase, MS-27-275, with marked in vivo antitumor activity against human tumors. | 1999 | Proc. Natl. Acad. Sci. U.S.A. | pmid:10200307 |
| Tavera-Mendoza LE et al. | Incorporation of histone deacetylase inhibition into the structure of a nuclear receptor agonist. | 2008 | Proc. Natl. Acad. Sci. U.S.A. | pmid:18550844 |
| Carmen AA et al. | Yeast HOS3 forms a novel trichostatin A-insensitive homodimer with intrinsic histone deacetylase activity. | 1999 | Proc. Natl. Acad. Sci. U.S.A. | pmid:10535926 |
| Monte M et al. | MAGE-A tumor antigens target p53 transactivation function through histone deacetylase recruitment and confer resistance to chemotherapeutic agents. | 2006 | Proc. Natl. Acad. Sci. U.S.A. | pmid:16847267 |
| Furumai R et al. | Potent histone deacetylase inhibitors built from trichostatin A and cyclic tetrapeptide antibiotics including trapoxin. | 2001 | Proc. Natl. Acad. Sci. U.S.A. | pmid:11134513 |
| Subramanian C et al. | Ku70 acetylation mediates neuroblastoma cell death induced by histone deacetylase inhibitors. | 2005 | Proc. Natl. Acad. Sci. U.S.A. | pmid:15778293 |
| Li B et al. | FOXP3 interactions with histone acetyltransferase and class II histone deacetylases are required for repression. | 2007 | Proc. Natl. Acad. Sci. U.S.A. | pmid:17360565 |