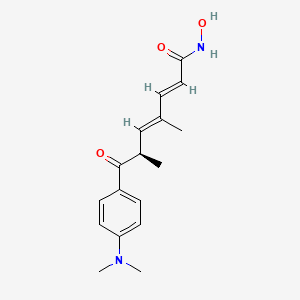
trichostatin A
Trichostatin is a lipid of Polyketides (PK) class. Trichostatin is associated with abnormalities such as Dentatorubral-Pallidoluysian Atrophy, PARAGANGLIOMAS 3, abnormal fragmented structure, Disintegration (morphologic abnormality) and Hyperostosis, Diffuse Idiopathic Skeletal. The involved functions are known as Acetylation, Cell Differentiation process, histone modification, Gene Silencing and Transcriptional Activation. Trichostatin often locates in CD41a, Hematopoietic System, Chromatin Structure, Blood and Endothelium. The associated genes with Trichostatin are SPI1 gene, CELL Gene, Chromatin, CXCR4 gene and DNMT1 gene. The related lipids are Butyrates, Promega, butyrate, Lipopolysaccharides and Steroids. The related experimental models are Knock-out, Mouse Model, Xenograft Model and Cancer Model.
Cross Reference
Introduction
To understand associated biological information of trichostatin A, we collected biological information of abnormalities, associated pathways, cellular/molecular locations, biological functions, related genes/proteins, lipids and common seen animal/experimental models with organized paragraphs from literatures.
What diseases are associated with trichostatin A?
trichostatin A is suspected in Infection, Morphologically altered structure, Ureteral obstruction, Photosensitization, Atherosclerosis, Hypertrophic Cardiomyopathy and other diseases in descending order of the highest number of associated sentences.
Related references are mostly published in these journals:
| Disease | Cross reference | Weighted score | Related literature |
|---|
Possible diseases from mapped MeSH terms on references
We collected disease MeSH terms mapped to the references associated with trichostatin A
PubChem Associated disorders and diseases
What pathways are associated with trichostatin A
Lipid pathways are not clear in current pathway databases. We organized associated pathways with trichostatin A through full-text articles, including metabolic pathways or pathways of biological mechanisms.
Related references are published most in these journals:
| Pathway name | Related literatures |
|---|
PubChem Biomolecular Interactions and Pathways
Link to PubChem Biomolecular Interactions and PathwaysWhat cellular locations are associated with trichostatin A?
Visualization in cellular structure
Associated locations are in red color. Not associated locations are in black.
Related references are published most in these journals:
| Location | Cross reference | Weighted score | Related literatures |
|---|
What functions are associated with trichostatin A?
Related references are published most in these journals:
| Function | Cross reference | Weighted score | Related literatures |
|---|
What lipids are associated with trichostatin A?
Related references are published most in these journals:
| Lipid concept | Cross reference | Weighted score | Related literatures |
|---|
What genes are associated with trichostatin A?
Related references are published most in these journals:
| Gene | Cross reference | Weighted score | Related literatures |
|---|
What common seen animal models are associated with trichostatin A?
Mouse Model
Mouse Model are used in the study 'Regulation of minichromosome maintenance gene family by microRNA-1296 and genistein in prostate cancer.' (Majid S et al., 2010), Mouse Model are used in the study 'Reversal of hypermethylation and reactivation of p16INK4a, RARbeta, and MGMT genes by genistein and other isoflavones from soy.' (Fang MZ et al., 2005) and Mouse Model are used in the study 'Histone deacetylase 3 mediates allergic skin inflammation by regulating expression of MCP1 protein.' (Kim Y et al., 2012).
Xenograft Model
Xenograft Model are used in the study 'Histone deacetylase inhibitors induce growth arrest and differentiation in uveal melanoma.' (Landreville S et al., 2012), Xenograft Model are used in the study 'Extended treatment with physiologic concentrations of dietary phytochemicals results in altered gene expression, reduced growth, and apoptosis of cancer cells.' (Moiseeva EP et al., 2007) and Xenograft Model are used in the study 'Retinoic acid and the histone deacetylase inhibitor trichostatin a inhibit the proliferation of human renal cell carcinoma in a xenograft tumor model.' (Touma SE et al., 2005).
Cancer Model
Cancer Model are used in the study 'Plasma pharmacokinetics and metabolism of the histone deacetylase inhibitor trichostatin a after intraperitoneal administration to mice.' (Sanderson L et al., 2004).
Related references are published most in these journals:
| Model | Cross reference | Weighted score | Related literatures |
|---|
NCBI Entrez Crosslinks
All references with trichostatin A
Download all related citations| Authors | Title | Published | Journal | PubMed Link |
|---|---|---|---|---|
| Chik CL et al. | Histone modifications on the adrenergic induction of type II deiodinase in rat pinealocytes. | 2011 | Mol. Cell. Endocrinol. | pmid:21704117 |
| Kanasaki H et al. | Trichostatin A reduces GnRH mRNA expression with a concomitant increase in retinaldehyde dehydrogenase in GnRH-producing neurons. | 2015 | Mol. Cell. Endocrinol. | pmid:26116234 |
| Goeman F et al. | Growth inhibition by the tumor suppressor p33ING1 in immortalized and primary cells: involvement of two silencing domains and effect of Ras. | 2005 | Mol. Cell. Biol. | pmid:15601862 |
| Abderrahmani A et al. | The transcriptional repressor REST determines the cell-specific expression of the human MAPK8IP1 gene encoding IB1 (JIP-1). | 2001 | Mol. Cell. Biol. | pmid:11585908 |
| Ashburner BP et al. | The p65 (RelA) subunit of NF-kappaB interacts with the histone deacetylase (HDAC) corepressors HDAC1 and HDAC2 to negatively regulate gene expression. | 2001 | Mol. Cell. Biol. | pmid:11564889 |
| Amann JM et al. | ETO, a target of t(8;21) in acute leukemia, makes distinct contacts with multiple histone deacetylases and binds mSin3A through its oligomerization domain. | 2001 | Mol. Cell. Biol. | pmid:11533236 |
| Plant KE et al. | Intergenic transcription in the human beta-globin gene cluster. | 2001 | Mol. Cell. Biol. | pmid:11533239 |
| Fang S et al. | Coordinated recruitment of histone methyltransferase G9a and other chromatin-modifying enzymes in SHP-mediated regulation of hepatic bile acid metabolism. | 2007 | Mol. Cell. Biol. | pmid:17145766 |
| Solomon JM et al. | Inhibition of SIRT1 catalytic activity increases p53 acetylation but does not alter cell survival following DNA damage. | 2006 | Mol. Cell. Biol. | pmid:16354677 |
| Kondo Y et al. | Critical role of histone methylation in tumor suppressor gene silencing in colorectal cancer. | 2003 | Mol. Cell. Biol. | pmid:12482974 |
| Ding Z et al. | Human MI-ER1 alpha and beta function as transcriptional repressors by recruitment of histone deacetylase 1 to their conserved ELM2 domain. | 2003 | Mol. Cell. Biol. | pmid:12482978 |
| Hwang CK et al. | Evidence of endogenous mu opioid receptor regulation by epigenetic control of the promoters. | 2007 | Mol. Cell. Biol. | pmid:17452465 |
| Zhang R et al. | HP1 proteins are essential for a dynamic nuclear response that rescues the function of perturbed heterochromatin in primary human cells. | 2007 | Mol. Cell. Biol. | pmid:17101789 |
| Bishop CL et al. | Role for centromeric heterochromatin and PML nuclear bodies in the cellular response to foreign DNA. | 2006 | Mol. Cell. Biol. | pmid:16537904 |
| Li J et al. | Transcriptional induction of MKP-1 in response to stress is associated with histone H3 phosphorylation-acetylation. | 2001 | Mol. Cell. Biol. | pmid:11689710 |
| Pivot-Pajot C et al. | Acetylation-dependent chromatin reorganization by BRDT, a testis-specific bromodomain-containing protein. | 2003 | Mol. Cell. Biol. | pmid:12861021 |
| Viollet B et al. | Embryonic but not postnatal reexpression of hepatocyte nuclear factor 1alpha (HNF1alpha) can reactivate the silent phenylalanine hydroxylase gene in HNF1alpha-deficient hepatocytes. | 2001 | Mol. Cell. Biol. | pmid:11340160 |
| Sutcliffe EL et al. | Dynamic histone variant exchange accompanies gene induction in T cells. | 2009 | Mol. Cell. Biol. | pmid:19158270 |
| Nusinzon I and Horvath CM | Positive and negative regulation of the innate antiviral response and beta interferon gene expression by deacetylation. | 2006 | Mol. Cell. Biol. | pmid:16581785 |
| Feng YQ et al. | Position effects are influenced by the orientation of a transgene with respect to flanking chromatin. | 2001 | Mol. Cell. Biol. | pmid:11113204 |
| Osborne A et al. | Histone deacetylase activity represses gamma interferon-inducible HLA-DR gene expression following the establishment of a DNase I-hypersensitive chromatin conformation. | 2001 | Mol. Cell. Biol. | pmid:11533238 |
| Sutcliffe JE et al. | Retinoblastoma protein disrupts interactions required for RNA polymerase III transcription. | 2000 | Mol. Cell. Biol. | pmid:11094071 |
| Zhang Y et al. | Coordinated changes in DNA methylation and histone modifications regulate silencing/derepression of luteinizing hormone receptor gene transcription. | 2005 | Mol. Cell. Biol. | pmid:16135786 |
| Hemavathy K et al. | Human Slug is a repressor that localizes to sites of active transcription. | 2000 | Mol. Cell. Biol. | pmid:10866665 |
| Curradi M et al. | Molecular mechanisms of gene silencing mediated by DNA methylation. | 2002 | Mol. Cell. Biol. | pmid:11940673 |
| Chen WY et al. | Histone deacetylase inhibitors reduce steroidogenesis through SCF-mediated ubiquitination and degradation of steroidogenic factor 1 (NR5A1). | 2007 | Mol. Cell. Biol. | pmid:17709382 |
| D'Alessio AC et al. | Acetylation-induced transcription is required for active DNA demethylation in methylation-silenced genes. | 2007 | Mol. Cell. Biol. | pmid:17709385 |
| Strouboulis J et al. | Transcriptional repression by XPc1, a new Polycomb homolog in Xenopus laevis embryos, is independent of histone deacetylase. | 1999 | Mol. Cell. Biol. | pmid:10330136 |
| DiRenzo J et al. | BRG-1 is recruited to estrogen-responsive promoters and cooperates with factors involved in histone acetylation. | 2000 | Mol. Cell. Biol. | pmid:11003650 |
| Madisen L et al. | The immunoglobulin heavy chain locus control region increases histone acetylation along linked c-myc genes. | 1998 | Mol. Cell. Biol. | pmid:9774645 |
| Adam E et al. | Potentiation of tumor necrosis factor-induced NF-kappa B activation by deacetylase inhibitors is associated with a delayed cytoplasmic reappearance of I kappa B alpha. | 2003 | Mol. Cell. Biol. | pmid:12917341 |
| Karki P et al. | Yin Yang 1 is a repressor of glutamate transporter EAAT2, and it mediates manganese-induced decrease of EAAT2 expression in astrocytes. | 2014 | Mol. Cell. Biol. | pmid:24469401 |
| Feng YQ et al. | Enhancer-dependent transcriptional oscillations in mouse erythroleukemia cells. | 1999 | Mol. Cell. Biol. | pmid:10373540 |
| Choy JS and Kron SJ | NuA4 subunit Yng2 function in intra-S-phase DNA damage response. | 2002 | Mol. Cell. Biol. | pmid:12417725 |
| Ghoshal K et al. | Inhibitors of histone deacetylase and DNA methyltransferase synergistically activate the methylated metallothionein I promoter by activating the transcription factor MTF-1 and forming an open chromatin structure. | 2002 | Mol. Cell. Biol. | pmid:12417732 |
| Hauser C et al. | Activation of the mouse histone deacetylase 1 gene by cooperative histone phosphorylation and acetylation. | 2002 | Mol. Cell. Biol. | pmid:12391151 |
| Westendorf JJ et al. | Runx2 (Cbfa1, AML-3) interacts with histone deacetylase 6 and represses the p21(CIP1/WAF1) promoter. | 2002 | Mol. Cell. Biol. | pmid:12391164 |
| Siddiqui H et al. | Histone deacetylation of RB-responsive promoters: requisite for specific gene repression but dispensable for cell cycle inhibition. | 2003 | Mol. Cell. Biol. | pmid:14560017 |
| Shetty S et al. | Transcription factor NF-kappaB differentially regulates death receptor 5 expression involving histone deacetylase 1. | 2005 | Mol. Cell. Biol. | pmid:15964798 |
| Kim HB et al. | Lipin 1 represses NFATc4 transcriptional activity in adipocytes to inhibit secretion of inflammatory factors. | 2010 | Mol. Cell. Biol. | pmid:20385772 |
| Dang VD et al. | A new member of the Sin3 family of corepressors is essential for cell viability and required for retroelement propagation in fission yeast. | 1999 | Mol. Cell. Biol. | pmid:10022921 |
| Ogryzko VV et al. | Human fibroblast commitment to a senescence-like state in response to histone deacetylase inhibitors is cell cycle dependent. | 1996 | Mol. Cell. Biol. | pmid:8756678 |
| Chang YC et al. | Cooperation of E2F-p130 and Sp1-pRb complexes in repression of the Chinese hamster dhfr gene. | 2001 | Mol. Cell. Biol. | pmid:11158299 |
| Zika E et al. | Histone deacetylase 1/mSin3A disrupts gamma interferon-induced CIITA function and major histocompatibility complex class II enhanceosome formation. | 2003 | Mol. Cell. Biol. | pmid:12697811 |
| Obrdlik A et al. | The histone acetyltransferase PCAF associates with actin and hnRNP U for RNA polymerase II transcription. | 2008 | Mol. Cell. Biol. | pmid:18710935 |
| Mihaylova VT et al. | Decreased expression of the DNA mismatch repair gene Mlh1 under hypoxic stress in mammalian cells. | 2003 | Mol. Cell. Biol. | pmid:12697826 |
| Kuwahara K et al. | The neuron-restrictive silencer element-neuron-restrictive silencer factor system regulates basal and endothelin 1-inducible atrial natriuretic peptide gene expression in ventricular myocytes. | 2001 | Mol. Cell. Biol. | pmid:11238943 |
| Jedrusik MA and Schulze E | Telomeric position effect variegation in Saccharomyces cerevisiae by Caenorhabditis elegans linker histones suggests a mechanistic connection between germ line and telomeric silencing. | 2003 | Mol. Cell. Biol. | pmid:12724425 |
| Naruse Y et al. | Circadian and light-induced transcription of clock gene Per1 depends on histone acetylation and deacetylation. | 2004 | Mol. Cell. Biol. | pmid:15226430 |
| Barr H et al. | Mbd2 contributes to DNA methylation-directed repression of the Xist gene. | 2007 | Mol. Cell. Biol. | pmid:17353271 |