| MeSH term | MeSH ID | Detail |
|---|---|---|
| Gestational Trophoblastic Disease | D031901 | 1 associated lipids |
| Carcinoma, Papillary, Follicular | D018265 | 1 associated lipids |
| Rubinstein-Taybi Syndrome | D012415 | 1 associated lipids |
| Lupus Vulgaris | D008177 | 1 associated lipids |
| Adenomyosis | D062788 | 1 associated lipids |
| Capsule Opacification | D058442 | 1 associated lipids |
| Goldenhar Syndrome | D006053 | 1 associated lipids |
| Classical Lissencephalies and Subcortical Band Heterotopias | D054221 | 1 associated lipids |
| Cystadenoma, Serous | D018293 | 1 associated lipids |
| Neoplasm Micrometastasis | D061206 | 1 associated lipids |
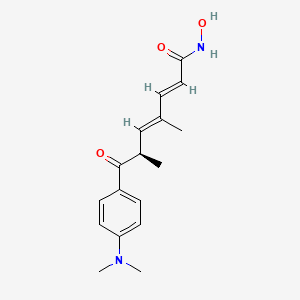
trichostatin A
Trichostatin is a lipid of Polyketides (PK) class. Trichostatin is associated with abnormalities such as Dentatorubral-Pallidoluysian Atrophy, PARAGANGLIOMAS 3, abnormal fragmented structure, Disintegration (morphologic abnormality) and Hyperostosis, Diffuse Idiopathic Skeletal. The involved functions are known as Acetylation, Cell Differentiation process, histone modification, Gene Silencing and Transcriptional Activation. Trichostatin often locates in CD41a, Hematopoietic System, Chromatin Structure, Blood and Endothelium. The associated genes with Trichostatin are SPI1 gene, CELL Gene, Chromatin, CXCR4 gene and DNMT1 gene. The related lipids are Butyrates, Promega, butyrate, Lipopolysaccharides and Steroids. The related experimental models are Knock-out, Mouse Model, Xenograft Model and Cancer Model.
Cross Reference
Introduction
To understand associated biological information of trichostatin A, we collected biological information of abnormalities, associated pathways, cellular/molecular locations, biological functions, related genes/proteins, lipids and common seen animal/experimental models with organized paragraphs from literatures.
What diseases are associated with trichostatin A?
trichostatin A is suspected in Infection, Morphologically altered structure, Ureteral obstruction, Photosensitization, Atherosclerosis, Hypertrophic Cardiomyopathy and other diseases in descending order of the highest number of associated sentences.
Related references are mostly published in these journals:
| Disease | Cross reference | Weighted score | Related literature |
|---|
Possible diseases from mapped MeSH terms on references
We collected disease MeSH terms mapped to the references associated with trichostatin A
PubChem Associated disorders and diseases
What pathways are associated with trichostatin A
Lipid pathways are not clear in current pathway databases. We organized associated pathways with trichostatin A through full-text articles, including metabolic pathways or pathways of biological mechanisms.
Related references are published most in these journals:
| Pathway name | Related literatures |
|---|
PubChem Biomolecular Interactions and Pathways
Link to PubChem Biomolecular Interactions and PathwaysWhat cellular locations are associated with trichostatin A?
Visualization in cellular structure
Associated locations are in red color. Not associated locations are in black.
Related references are published most in these journals:
| Location | Cross reference | Weighted score | Related literatures |
|---|
What functions are associated with trichostatin A?
Related references are published most in these journals:
| Function | Cross reference | Weighted score | Related literatures |
|---|
What lipids are associated with trichostatin A?
Related references are published most in these journals:
| Lipid concept | Cross reference | Weighted score | Related literatures |
|---|
What genes are associated with trichostatin A?
Related references are published most in these journals:
| Gene | Cross reference | Weighted score | Related literatures |
|---|
What common seen animal models are associated with trichostatin A?
Mouse Model
Mouse Model are used in the study 'Regulation of minichromosome maintenance gene family by microRNA-1296 and genistein in prostate cancer.' (Majid S et al., 2010), Mouse Model are used in the study 'Reversal of hypermethylation and reactivation of p16INK4a, RARbeta, and MGMT genes by genistein and other isoflavones from soy.' (Fang MZ et al., 2005) and Mouse Model are used in the study 'Histone deacetylase 3 mediates allergic skin inflammation by regulating expression of MCP1 protein.' (Kim Y et al., 2012).
Xenograft Model
Xenograft Model are used in the study 'Histone deacetylase inhibitors induce growth arrest and differentiation in uveal melanoma.' (Landreville S et al., 2012), Xenograft Model are used in the study 'Extended treatment with physiologic concentrations of dietary phytochemicals results in altered gene expression, reduced growth, and apoptosis of cancer cells.' (Moiseeva EP et al., 2007) and Xenograft Model are used in the study 'Retinoic acid and the histone deacetylase inhibitor trichostatin a inhibit the proliferation of human renal cell carcinoma in a xenograft tumor model.' (Touma SE et al., 2005).
Cancer Model
Cancer Model are used in the study 'Plasma pharmacokinetics and metabolism of the histone deacetylase inhibitor trichostatin a after intraperitoneal administration to mice.' (Sanderson L et al., 2004).
Related references are published most in these journals:
| Model | Cross reference | Weighted score | Related literatures |
|---|
NCBI Entrez Crosslinks
All references with trichostatin A
Download all related citations| Authors | Title | Published | Journal | PubMed Link |
|---|---|---|---|---|
| McBain JA et al. | Apoptotic death in adenocarcinoma cell lines induced by butyrate and other histone deacetylase inhibitors. | 1997 | Biochem. Pharmacol. | pmid:9214697 |
| Ohno Y et al. | Macrophage inflammatory protein-2: chromosomal regulation in rat small intestinal epithelial cells. | 1997 | Proc. Natl. Acad. Sci. U.S.A. | pmid:9294201 |
| Chen WY et al. | Reactivation of silenced, virally transduced genes by inhibitors of histone deacetylase. | 1997 | Proc. Natl. Acad. Sci. U.S.A. | pmid:9159154 |
| Minucci S et al. | A histone deacetylase inhibitor potentiates retinoid receptor action in embryonal carcinoma cells. | 1997 | Proc. Natl. Acad. Sci. U.S.A. | pmid:9326603 |
| Ekwall K et al. | Transient inhibition of histone deacetylation alters the structural and functional imprint at fission yeast centromeres. | 1997 | Cell | pmid:9428524 |
| Xu L et al. | Effect of the histone deacetylase inhibitor trichostatin A on the responsiveness of rat hepatocytes to dioxin. | 1997 | Biochem. Pharmacol. | pmid:9174108 |
| Stein P et al. | Stage-dependent redistributions of acetylated histones in nuclei of the early preimplantation mouse embryo. | 1997 | Mol. Reprod. Dev. | pmid:9211426 |
| Sommer A et al. | Cell growth inhibition by the Mad/Max complex through recruitment of histone deacetylase activity. | 1997 | Curr. Biol. | pmid:9197243 |
| Chen ZJ and Pikaard CS | Epigenetic silencing of RNA polymerase I transcription: a role for DNA methylation and histone modification in nucleolar dominance. | 1997 | Genes Dev. | pmid:9284051 |
| Medina V et al. | Induction of caspase-3 protease activity and apoptosis by butyrate and trichostatin A (inhibitors of histone deacetylase): dependence on protein synthesis and synergy with a mitochondrial/cytochrome c-dependent pathway. | 1997 | Cancer Res. | pmid:9288776 |
| Dangond F and Gullans SR | Differential expression of human histone deacetylase mRNAs in response to immune cell apoptosis induction by trichostatin A and butyrate. | 1998 | Biochem. Biophys. Res. Commun. | pmid:9647779 |
| Lin RJ et al. | Role of the histone deacetylase complex in acute promyelocytic leukaemia. | 1998 | Nature | pmid:9486654 |
| Grignani F et al. | Fusion proteins of the retinoic acid receptor-alpha recruit histone deacetylase in promyelocytic leukaemia. | 1998 | Nature | pmid:9486655 |
| Wong J et al. | Distinct requirements for chromatin assembly in transcriptional repression by thyroid hormone receptor and histone deacetylase. | 1998 | EMBO J. | pmid:9430643 |
| Komatsu Y and Hayashi H | Histone deacetylase inhibitors up-regulate the expression of cell surface MHC class-I molecules in B16/BL6 cells. | 1998 | J. Antibiot. | pmid:9531994 |
| Magnaghi-Jaulin L et al. | Retinoblastoma protein represses transcription by recruiting a histone deacetylase. | 1998 | Nature | pmid:9468140 |
| Svensson K et al. | The paternal allele of the H19 gene is progressively silenced during early mouse development: the acetylation status of histones may be involved in the generation of variegated expression patterns. | 1998 | Development | pmid:9389664 |
| Emiliani S et al. | Characterization of a human RPD3 ortholog, HDAC3. | 1998 | Proc. Natl. Acad. Sci. U.S.A. | pmid:9501169 |
| El Kharroubi A et al. | Transcriptional activation of the integrated chromatin-associated human immunodeficiency virus type 1 promoter. | 1998 | Mol. Cell. Biol. | pmid:9566873 |
| Gray SG and Ekström TJ | Effects of cell density and trichostatin A on the expression of HDAC1 and p57Kip2 in Hep 3B cells. | 1998 | Biochem. Biophys. Res. Commun. | pmid:9571167 |
| Belyaev ND et al. | The acetylation patterns of histones H3 and H4 along Vicia faba chromosomes are different. | 1998 | Chromosome Res. | pmid:9510512 |
| Nan X et al. | Transcriptional repression by the methyl-CpG-binding protein MeCP2 involves a histone deacetylase complex. | 1998 | Nature | pmid:9620804 |
| Kim YB et al. | phd1+, a histone deacetylase gene of Schizosaccharomyces pombe, is required for the meiotic cell cycle and resistance to trichostatin A. | 1998 | FEBS Lett. | pmid:9781677 |
| Richon VM et al. | A class of hybrid polar inducers of transformed cell differentiation inhibits histone deacetylases. | 1998 | Proc. Natl. Acad. Sci. U.S.A. | pmid:9501205 |
| Egawa K et al. | Identification of active substances from Streptomyces culture fluids using p53-independent expression of p21/WAF1/Cip1 gene and their mode of action. | 1998 | Biol. Pharm. Bull. | pmid:9781835 |
| Kohge T et al. | Promotion of antigen-specific antibody production in murine B cells by a moderate increase in histone acetylation. | 1998 | Biochem. Pharmacol. | pmid:9825735 |
| Grewal SI et al. | Histone deacetylase homologs regulate epigenetic inheritance of transcriptional silencing and chromosome segregation in fission yeast. | 1998 | Genetics | pmid:9755190 |
| Preston CM and McFarlane M | Cytodifferentiating agents affect the replication of herpes simplex virus type 1 in the absence of functional VP16. | 1998 | Virology | pmid:9791032 |
| Hu JF et al. | The role of histone acetylation in the allelic expression of the imprinted human insulin-like growth factor II gene. | 1998 | Biochem. Biophys. Res. Commun. | pmid:9792787 |
| Llorente A et al. | Expression of mutant dynamin inhibits toxicity and transport of endocytosed ricin to the Golgi apparatus. | 1998 | J. Cell Biol. | pmid:9456316 |
| Korhonen P et al. | Expression of transcriptional repressor protein mSin3A but not mSin3B is induced during neuronal apoptosis. | 1998 | Biochem. Biophys. Res. Commun. | pmid:9813182 |
| Waterborg JH | Dynamics of histone acetylation in Chlamydomonas reinhardtii. | 1998 | J. Biol. Chem. | pmid:9765294 |
| Madisen L et al. | The immunoglobulin heavy chain locus control region increases histone acetylation along linked c-myc genes. | 1998 | Mol. Cell. Biol. | pmid:9774645 |
| Eden S et al. | DNA methylation models histone acetylation. | 1998 | Nature | pmid:9732866 |
| Phelan MW et al. | Hypoxia increases thrombospondin-1 transcript and protein in cultured endothelial cells. | 1998 | J. Lab. Clin. Med. | pmid:9851743 |
| Ciana P et al. | Leukemic transformation by the v-ErbA oncoprotein entails constitutive binding to and repression of an erythroid enhancer in vivo. | 1998 | EMBO J. | pmid:9857194 |
| De Luca P et al. | Retinoblastoma protein tethered to promoter DNA represses TBP-mediated transcription. | 1998 | J. Cell. Biochem. | pmid:9671233 |
| Selker EU | Trichostatin A causes selective loss of DNA methylation in Neurospora. | 1998 | Proc. Natl. Acad. Sci. U.S.A. | pmid:9689097 |
| Jin S and Scotto KW | Transcriptional regulation of the MDR1 gene by histone acetyltransferase and deacetylase is mediated by NF-Y. | 1998 | Mol. Cell. Biol. | pmid:9632821 |
| Nakajima H et al. | FR901228, a potent antitumor antibiotic, is a novel histone deacetylase inhibitor. | 1998 | Exp. Cell Res. | pmid:9633520 |
| Thompson EM and Renard JP | Preferential nuclear location of a transgene does not depend on its transcriptional activity during early mouse development. | 1998 | Chromosoma | pmid:9880765 |
| Durum SK et al. | Interleukin 7 receptor control of T cell receptor gamma gene rearrangement: role of receptor-associated chains and locus accessibility. | 1998 | J. Exp. Med. | pmid:9858510 |
| Nemer M | Histone deacetylase mRNA temporally and spatially regulated in its expression in sea urchin embryos. | 1998 | Dev. Growth Differ. | pmid:9865968 |
| Ishiguro K and Sartorelli AC | Coinduction of embryonic and adult-type globin mRNAs by sodium butyrate and trichostatin A in two murine interleukin-3-dependent bone marrow-derived cell lines. | 1998 | Blood | pmid:9834245 |
| Walia H et al. | Histone acetylation is required to maintain the unfolded nucleosome structure associated with transcribing DNA. | 1998 | J. Biol. Chem. | pmid:9603965 |
| Taddei A et al. | Duplication and maintenance of heterochromatin domains. | 1999 | J. Cell Biol. | pmid:10601331 |
| Ghosh AK et al. | MBP-1 physically associates with histone deacetylase for transcriptional repression. | 1999 | Biochem. Biophys. Res. Commun. | pmid:10403782 |
| Gradin K et al. | Repression of dioxin signal transduction in fibroblasts. Identification Of a putative repressor associated with Arnt. | 1999 | J. Biol. Chem. | pmid:10224119 |
| Liu Z et al. | Steroid receptor coactivator-1 (SRC-1) enhances ligand-dependent and receptor-dependent cell-free transcription of chromatin. | 1999 | Proc. Natl. Acad. Sci. U.S.A. | pmid:10449719 |
| Wang J et al. | Inhibitors of histone deacetylase relieve ETO-mediated repression and induce differentiation of AML1-ETO leukemia cells. | 1999 | Cancer Res. | pmid:10383127 |