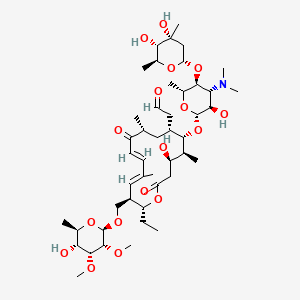
Tylosin
Tylosin is a lipid of Polyketides (PK) class. Tylosin is associated with abnormalities such as Mastitis, Bovine, Infection, Bacterial Infections, Arthritis and Ileitis. The involved functions are known as Anabolism, acireductone dioxygenase [iron(II)-requiring] activity, Protein Biosynthesis, Mastitis and Methylation. Tylosin often locates in Ribosomes, Cell Wall, 50S ribosomal subunit, Ribosome Subunits, Large and Ribosome Subunits. The associated genes with Tylosin are Gene Clusters, Genome, resistance genes, Homologous Gene and HM13 gene. The related experimental models are Knock-out.
Cross Reference
Introduction
To understand associated biological information of Tylosin, we collected biological information of abnormalities, associated pathways, cellular/molecular locations, biological functions, related genes/proteins, lipids and common seen animal/experimental models with organized paragraphs from literatures.
What diseases are associated with Tylosin?
Tylosin is suspected in Porcine ulcerative spirochetosis, Infection, Liver Abscess, Respiration Disorders, Intestinal Diseases, Mastitis, Bovine and other diseases in descending order of the highest number of associated sentences.
Related references are mostly published in these journals:
- Antimicrob. Agents Chemother. (6)
- Appl. Environ. Microbiol. (2)
- Proc. Natl. Acad. Sci. U.S.A. (2)
- Others (6)
| Disease | Cross reference | Weighted score | Related literature |
|---|
Possible diseases from mapped MeSH terms on references
We collected disease MeSH terms mapped to the references associated with Tylosin
PubChem Associated disorders and diseases
What pathways are associated with Tylosin
There are no associated biomedical information in the current reference collection.
PubChem Biomolecular Interactions and Pathways
Link to PubChem Biomolecular Interactions and PathwaysWhat cellular locations are associated with Tylosin?
Visualization in cellular structure
Associated locations are in red color. Not associated locations are in black.
Related references are published most in these journals:
| Location | Cross reference | Weighted score | Related literatures |
|---|
What functions are associated with Tylosin?
Related references are published most in these journals:
| Function | Cross reference | Weighted score | Related literatures |
|---|
What lipids are associated with Tylosin?
There are no associated biomedical information in the current reference collection.
What genes are associated with Tylosin?
Related references are published most in these journals:
- Antimicrob. Agents Chemother. (9)
- Microbiology (Reading, Engl.) (3)
- Proc. Natl. Acad. Sci. U.S.A. (2)
- Others (13)
| Gene | Cross reference | Weighted score | Related literatures |
|---|
What common seen animal models are associated with Tylosin?
Knock-out
Knock-out are used in the study 'A second tylosin resistance determinant, Erm B, in Arcanobacterium pyogenes.' (Jost BH et al., 2004).
Related references are published most in these journals:
| Model | Cross reference | Weighted score | Related literatures |
|---|
NCBI Entrez Crosslinks
All references with Tylosin
Download all related citations| Authors | Title | Published | Journal | PubMed Link |
|---|---|---|---|---|
| Pérez RA et al. | Analysis of macrolide antibiotics in water by magnetic solid-phase extraction and liquid chromatography-tandem mass spectrometry. | 2017 | J Pharm Biomed Anal | pmid:28858671 |
| Sato T et al. | Mycoplasma bovis isolates from dairy calves in Japan have less susceptibility than a reference strain to all approved macrolides associated with a point mutation (G748A) combined with multiple species-specific nucleotide alterations in 23S rRNA. | 2017 | Microbiol. Immunol. | pmid:28504455 |
| Nowakiewicz A et al. | Determination of antimicrobial resistance of Enterococcus strains isolated from pigs and their genotypic characterization by method of amplification of DNA fragments surrounding rare restriction sites (ADSRRS fingerprinting). | 2017 | J. Med. Microbiol. | pmid:28260584 |
| Hossain A et al. | Occurrence, distribution, ecological and resistance risks of antibiotics in surface water of finfish and shellfish aquaculture in Bangladesh. | 2017 | Chemosphere | pmid:28888121 |
| Amachawadi RG et al. | Bacterial flora of liver abscesses in crossbred beef cattle and Holstein steers fed finishing diets with or without tylosin. | 2017 | J. Anim. Sci. | pmid:28805921 |
| Sugawara A et al. | 5-O-Mycaminosyltylonolide antibacterial derivatives: design, synthesis and bioactivity. | 2017 | J. Antibiot. | pmid:28559578 |
| Abell KM et al. | A mixed treatment comparison meta-analysis of metaphylaxis treatments for bovine respiratory disease in beef cattle. | 2017 | J. Anim. Sci. | pmid:28380607 |
| Ray P et al. | Fate and Effect of Antibiotics in Beef and Dairy Manure during Static and Turned Composting. | 2017 | J. Environ. Qual. | pmid:28177414 |
| Liu B et al. | Copper exposure to soil under single and repeated application: Selection for the microbial community tolerance and effects on the dissipation of antibiotics. | 2017 | J. Hazard. Mater. | pmid:27930997 |
| Qin Y et al. | Analysis of sulfonamides, tilmicosin and avermectins residues in typical animal matrices with multi-plug filtration cleanup by liquid chromatography-tandem mass spectrometry detection. | 2017 | J. Chromatogr. B Analyt. Technol. Biomed. Life Sci. | pmid:28410479 |