| MeSH term | MeSH ID | Detail |
|---|---|---|
| Drug Hypersensitivity | D004342 | 20 associated lipids |
| Bacterial Infections | D001424 | 21 associated lipids |
| Body Weight | D001835 | 333 associated lipids |
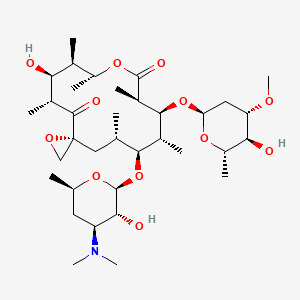
oleandomycin
oleandomycin is a lipid of Polyketides (PK) class. The involved functions are known as 5-(carboxyamino)imidazole ribonucleotide mutase activity, Methylation, enzyme activity, Anabolism and Biosynthetic Pathways. Oleandomycin often locates in Chromosomes and Protoplasm. The associated genes with oleandomycin are resistance genes, Open Reading Frames, CTTNBP2 gene, Gene Feature and Gene Clusters.
Cross Reference
Introduction
To understand associated biological information of oleandomycin, we collected biological information of abnormalities, associated pathways, cellular/molecular locations, biological functions, related genes/proteins, lipids and common seen animal/experimental models with organized paragraphs from literatures.
What diseases are associated with oleandomycin?
There are no associated biomedical information in the current reference collection.
Possible diseases from mapped MeSH terms on references
We collected disease MeSH terms mapped to the references associated with oleandomycin
PubChem Associated disorders and diseases
What pathways are associated with oleandomycin
There are no associated biomedical information in the current reference collection.
PubChem Biomolecular Interactions and Pathways
Link to PubChem Biomolecular Interactions and PathwaysWhat cellular locations are associated with oleandomycin?
Visualization in cellular structure
Associated locations are in red color. Not associated locations are in black.
Related references are published most in these journals:
| Location | Cross reference | Weighted score | Related literatures |
|---|
What functions are associated with oleandomycin?
Related references are published most in these journals:
| Function | Cross reference | Weighted score | Related literatures |
|---|
What lipids are associated with oleandomycin?
There are no associated biomedical information in the current reference collection.
What genes are associated with oleandomycin?
Related references are published most in these journals:
| Gene | Cross reference | Weighted score | Related literatures |
|---|
What common seen animal models are associated with oleandomycin?
There are no associated biomedical information in the current reference collection.
NCBI Entrez Crosslinks
All references with oleandomycin
Download all related citations| Authors | Title | Published | Journal | PubMed Link |
|---|---|---|---|---|
| Di Modugno V et al. | Low level resistance to oleandomycin as a marker of ermA in staphylococci. | 2002 | J. Antimicrob. Chemother. | pmid:11815596 |
| Smirnov VV et al. | [Gram-negative eubacteria isolated from the water, molluscs and algae of the Black Sea]. | 2001 Jul-Aug | Mikrobiol. Z. | pmid:11692675 |
| RodrÃguez L et al. | Functional analysis of OleY L-oleandrosyl 3-O-methyltransferase of the oleandomycin biosynthetic pathway in Streptomyces antibioticus. | 2001 | J. Bacteriol. | pmid:11514520 |
| Kimura Y et al. | Characterization of the mac-1 gene encoding a putative ABC transporter from Myxococcus xanthus. | 2001 | J. Biochem. | pmid:11226873 |
| Garzotti M et al. | Coupling of a supercritical fluid chromatography system to a hybrid (Q-TOF 2) mass spectrometer: on-line accurate mass measurements. | 2001 | Rapid Commun. Mass Spectrom. | pmid:11445901 |
| Quirós LM et al. | Glycosylation of macrolide antibiotics. Purification and kinetic studies of a macrolide glycosyltransferase from Streptomyces antibioticus. | 2000 | J. Biol. Chem. | pmid:10766792 |
| Aguirrezabalaga I et al. | Identification and expression of genes involved in biosynthesis of L-oleandrose and its intermediate L-olivose in the oleandomycin producer Streptomyces antibioticus. | 2000 | Antimicrob. Agents Chemother. | pmid:10770761 |
| Bajic S et al. | Analysis of erythromycin by liquid chromatography/mass spectrometry using involatile mobile phases with a novel atmospheric pressure ionization source. | 2000 | Rapid Commun. Mass Spectrom. | pmid:10637421 |
| Tang L et al. | Formation of functional heterologous complexes using subunits from the picromycin, erythromycin and oleandomycin polyketide synthases. | 2000 | Chem. Biol. | pmid:10662693 |
| Zhou J et al. | Separation and determination of the macrolide antibiotics (erythromycin, spiramycin and oleandomycin) by capillary electrophoresis coupled with fast reductive voltammetric detection. | 2000 | Electrophoresis | pmid:10826680 |