| MeSH term | MeSH ID | Detail |
|---|---|---|
| Drug Hypersensitivity | D004342 | 20 associated lipids |
| Bacterial Infections | D001424 | 21 associated lipids |
| Body Weight | D001835 | 333 associated lipids |
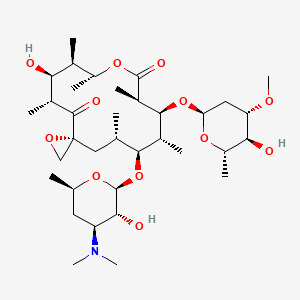
oleandomycin
oleandomycin is a lipid of Polyketides (PK) class. The involved functions are known as 5-(carboxyamino)imidazole ribonucleotide mutase activity, Methylation, enzyme activity, Anabolism and Biosynthetic Pathways. Oleandomycin often locates in Chromosomes and Protoplasm. The associated genes with oleandomycin are resistance genes, Open Reading Frames, CTTNBP2 gene, Gene Feature and Gene Clusters.
Cross Reference
Introduction
To understand associated biological information of oleandomycin, we collected biological information of abnormalities, associated pathways, cellular/molecular locations, biological functions, related genes/proteins, lipids and common seen animal/experimental models with organized paragraphs from literatures.
What diseases are associated with oleandomycin?
There are no associated biomedical information in the current reference collection.
Possible diseases from mapped MeSH terms on references
We collected disease MeSH terms mapped to the references associated with oleandomycin
PubChem Associated disorders and diseases
What pathways are associated with oleandomycin
There are no associated biomedical information in the current reference collection.
PubChem Biomolecular Interactions and Pathways
Link to PubChem Biomolecular Interactions and PathwaysWhat cellular locations are associated with oleandomycin?
Visualization in cellular structure
Associated locations are in red color. Not associated locations are in black.
Related references are published most in these journals:
| Location | Cross reference | Weighted score | Related literatures |
|---|
What functions are associated with oleandomycin?
Related references are published most in these journals:
| Function | Cross reference | Weighted score | Related literatures |
|---|
What lipids are associated with oleandomycin?
There are no associated biomedical information in the current reference collection.
What genes are associated with oleandomycin?
Related references are published most in these journals:
| Gene | Cross reference | Weighted score | Related literatures |
|---|
What common seen animal models are associated with oleandomycin?
There are no associated biomedical information in the current reference collection.
NCBI Entrez Crosslinks
All references with oleandomycin
Download all related citations| Authors | Title | Published | Journal | PubMed Link |
|---|---|---|---|---|
| Aguirrezabalaga I et al. | Identification and expression of genes involved in biosynthesis of L-oleandrose and its intermediate L-olivose in the oleandomycin producer Streptomyces antibioticus. | 2000 | Antimicrob. Agents Chemother. | pmid:10770761 |
| Rodriguez L et al. | Generation of hybrid elloramycin analogs by combinatorial biosynthesis using genes from anthracycline-type and macrolide biosynthetic pathways. | 2000 | J. Mol. Microbiol. Biotechnol. | pmid:10937435 |
| Shah S et al. | Cloning, characterization and heterologous expression of a polyketide synthase and P-450 oxidase involved in the biosynthesis of the antibiotic oleandomycin. | 2000 | J. Antibiot. | pmid:10908114 |
| Quirós LM et al. | Inversion of the anomeric configuration of the transferred sugar during inactivation of the macrolide antibiotic oleandomycin catalyzed by a macrolide glycosyltransferase. | 2000 | FEBS Lett. | pmid:10913610 |
| Zhou J et al. | Separation and determination of the macrolide antibiotics (erythromycin, spiramycin and oleandomycin) by capillary electrophoresis coupled with fast reductive voltammetric detection. | 2000 | Electrophoresis | pmid:10826680 |
| Doumith M et al. | Interspecies complementation in Saccharopolyspora erythraea : elucidation of the function of oleP1, oleG1 and oleG2 from the oleandomycin biosynthetic gene cluster of Streptomyces antibioticus and generation of new erythromycin derivatives. | 1999 | Mol. Microbiol. | pmid:10594828 |
| Taniguchi K et al. | Identification of functional amino acids in the macrolide 2'-phosphotransferase II. | 1999 | Antimicrob. Agents Chemother. | pmid:10428938 |
| Taniguchi K et al. | Identification of Escherichia coli clinical isolates producing macrolide 2'-phosphotransferase by a highly sensitive detection method. | 1998 | FEMS Microbiol. Lett. | pmid:9809420 |
| Quirós LM et al. | Two glycosyltransferases and a glycosidase are involved in oleandomycin modification during its biosynthesis by Streptomyces antibioticus. | 1998 | Mol. Microbiol. | pmid:9680207 |
| Hoeben D et al. | Antibiotics commonly used to treat mastitis and respiratory burst of bovine polymorphonuclear leukocytes. | 1998 | J. Dairy Sci. | pmid:9532493 |