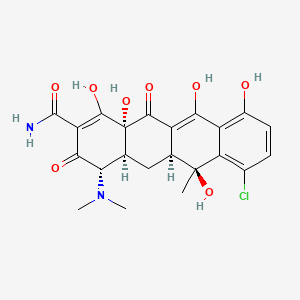
chlortetracycline
chlortetracycline is a lipid of Polyketides (PK) class. Chlortetracycline is associated with abnormalities such as Granulomatous Disease, Chronic, Infection, Ischemia, Cerebral Ischemia and Cerebral Infarction. The involved functions are known as Regulation, Binding (Molecular Function), Agent, Stimulus and Process. Chlortetracycline often locates in Protoplasm, Plasma membrane, Membrane, Cytoplasm and specific granule. The associated genes with chlortetracycline are FPR1 gene, P4HTM gene, Homologous Gene, HIST1H1C gene and Microbiome. The related lipids are Lysophosphatidylcholines, Sterols, dilauroyl lecithin, seminolipid and Total cholesterol. The related experimental models are Mouse Model.
Cross Reference
Introduction
To understand associated biological information of chlortetracycline, we collected biological information of abnormalities, associated pathways, cellular/molecular locations, biological functions, related genes/proteins, lipids and common seen animal/experimental models with organized paragraphs from literatures.
What diseases are associated with chlortetracycline?
chlortetracycline is suspected in Ischemia, Cerebral Ischemia, Cerebral Infarction, Granulomatous Disease, Chronic, Infection, Antibiotic resistant infection and other diseases in descending order of the highest number of associated sentences.
Related references are mostly published in these journals:
| Disease | Cross reference | Weighted score | Related literature |
|---|
Possible diseases from mapped MeSH terms on references
We collected disease MeSH terms mapped to the references associated with chlortetracycline
PubChem Associated disorders and diseases
What pathways are associated with chlortetracycline
There are no associated biomedical information in the current reference collection.
PubChem Biomolecular Interactions and Pathways
Link to PubChem Biomolecular Interactions and PathwaysWhat cellular locations are associated with chlortetracycline?
Visualization in cellular structure
Associated locations are in red color. Not associated locations are in black.
Related references are published most in these journals:
| Location | Cross reference | Weighted score | Related literatures |
|---|
What functions are associated with chlortetracycline?
Related references are published most in these journals:
| Function | Cross reference | Weighted score | Related literatures |
|---|
What lipids are associated with chlortetracycline?
Related references are published most in these journals:
| Lipid concept | Cross reference | Weighted score | Related literatures |
|---|
What genes are associated with chlortetracycline?
Related references are published most in these journals:
| Gene | Cross reference | Weighted score | Related literatures |
|---|
What common seen animal models are associated with chlortetracycline?
Mouse Model
Mouse Model are used in the study 'Chlortetracycline and demeclocycline inhibit calpains and protect mouse neurons against glutamate toxicity and cerebral ischemia.' (Jiang SX et al., 2005).
Related references are published most in these journals:
| Model | Cross reference | Weighted score | Related literatures |
|---|
NCBI Entrez Crosslinks
All references with chlortetracycline
Download all related citations| Authors | Title | Published | Journal | PubMed Link |
|---|---|---|---|---|
| You N et al. | Development and evaluation of diffusive gradients in thin films based on nano-sized zinc oxide particles for the in situ sampling of tetracyclines in pig breeding wastewater. | 2019 | Sci. Total Environ. | pmid:30312908 |
| Deitchman AN et al. | Enhanced in vitro activity of tigecycline in the presence of chelating agents. | 2018 | Int. J. Antimicrob. Agents | pmid:29305959 |
| Schwake-Anduschus C and Langenkämper G | Chlortetracycline and related tetracyclines: detection in wheat and rye grain. | 2018 | J. Sci. Food Agric. | pmid:29484666 |
| Wallace JS et al. | Occurrence and transformation of veterinary antibiotics and antibiotic resistance genes in dairy manure treated by advanced anaerobic digestion and conventional treatment methods. | 2018 | Environ. Pollut. | pmid:29455089 |
| Xiong W et al. | Antibiotic-mediated changes in the fecal microbiome of broiler chickens define the incidence of antibiotic resistance genes. | 2018 | Microbiome | pmid:29439741 |
| Yan H et al. | Changes to tetracyclines and tetracycline resistance genes in arable soils after single and multiple applications of manure containing tetracyclines. | 2018 | Environ Sci Pollut Res Int | pmid:29222656 |
| Pandey RP et al. | Bioconversion of Tetracycline Antibiotics to Novel Glucoside Derivatives by Single-Vessel Multienzymatic Glycosylation. | 2018 | J. Microbiol. Biotechnol. | pmid:29212298 |
| Yin F et al. | Antibiotic degradation and microbial community structures during acidification and methanogenesis of swine manure containing chlortetracycline or oxytetracycline. | 2018 | Bioresour. Technol. | pmid:29174902 |
| Zhang Z et al. | Highly luminescent nitrogen-doped carbon dots for simultaneous determination of chlortetracycline and sulfasalazine. | 2018 | Luminescence | pmid:29044942 |
| Li W et al. | Quantitative proteomic analysis reveals that chemotaxis is involved in chlortetracycline resistance of Aeromonas hydrophila. | 2018 | J Proteomics | pmid:28986269 |