| MeSH term | MeSH ID | Detail |
|---|---|---|
| Melanoma | D008545 | 69 associated lipids |
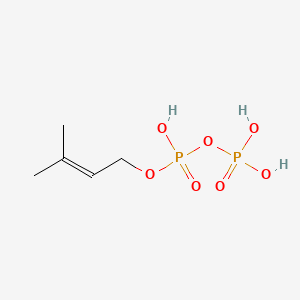
Dimethylallyl pyrophosphate
Dimethylallyl pyrophosphate is a lipid of Prenol Lipids (PR) class. Dimethylallyl pyrophosphate is associated with abnormalities such as Consumption-archaic term for TB and Wiskott-Aldrich Syndrome. The involved functions are known as Anabolism, Biochemical Pathway, Oxidation, Process and Chelating Activity [MoA]. Dimethylallyl pyrophosphate often locates in Chloroplasts, Plastids, chloroplast stroma, Cytosol and Cell membrane. The associated genes with Dimethylallyl pyrophosphate are IRF6 wt Allele and ADRBK1 gene. The related lipids are Sterols.
Cross Reference
Introduction
To understand associated biological information of Dimethylallyl pyrophosphate, we collected biological information of abnormalities, associated pathways, cellular/molecular locations, biological functions, related genes/proteins, lipids and common seen animal/experimental models with organized paragraphs from literatures.
What diseases are associated with Dimethylallyl pyrophosphate?
Dimethylallyl pyrophosphate is suspected in and other diseases in descending order of the highest number of associated sentences.
Related references are mostly published in these journals:
| Disease | Cross reference | Weighted score | Related literature |
|---|
Possible diseases from mapped MeSH terms on references
We collected disease MeSH terms mapped to the references associated with Dimethylallyl pyrophosphate
PubChem Associated disorders and diseases
What pathways are associated with Dimethylallyl pyrophosphate
Lipid pathways are not clear in current pathway databases. We organized associated pathways with Dimethylallyl pyrophosphate through full-text articles, including metabolic pathways or pathways of biological mechanisms.
Related references are published most in these journals:
| Pathway name | Related literatures |
|---|
PubChem Biomolecular Interactions and Pathways
Link to PubChem Biomolecular Interactions and PathwaysWhat cellular locations are associated with Dimethylallyl pyrophosphate?
Visualization in cellular structure
Associated locations are in red color. Not associated locations are in black.
Related references are published most in these journals:
| Location | Cross reference | Weighted score | Related literatures |
|---|
What functions are associated with Dimethylallyl pyrophosphate?
Related references are published most in these journals:
| Function | Cross reference | Weighted score | Related literatures |
|---|
What lipids are associated with Dimethylallyl pyrophosphate?
Related references are published most in these journals:
| Lipid concept | Cross reference | Weighted score | Related literatures |
|---|
What genes are associated with Dimethylallyl pyrophosphate?
Related references are published most in these journals:
| Gene | Cross reference | Weighted score | Related literatures |
|---|
What common seen animal models are associated with Dimethylallyl pyrophosphate?
There are no associated biomedical information in the current reference collection.
NCBI Entrez Crosslinks
All references with Dimethylallyl pyrophosphate
Download all related citations| Authors | Title | Published | Journal | PubMed Link |
|---|---|---|---|---|
| Fischer MJ et al. | Identification of a lysine residue important for the catalytic activity of yeast farnesyl diphosphate synthase. | 2011 | Protein J. | pmid:21643844 |
| Hunter WN | Isoprenoid precursor biosynthesis offers potential targets for drug discovery against diseases caused by apicomplexan parasites. | 2011 | Curr Top Med Chem | pmid:21619509 |
| Hemmi H | [Unique flavoenzyme: type 2 isopentenyl diphosphate isomerase]. | 2011 | Seikagaku | pmid:21626882 |
| Thabet I et al. | The subcellular localization of periwinkle farnesyl diphosphate synthase provides insight into the role of peroxisome in isoprenoid biosynthesis. | 2011 | J. Plant Physiol. | pmid:21872968 |
| Wen W and Yu R | Artemisinin biosynthesis and its regulatory enzymes: Progress and perspective. | 2011 | Pharmacogn Rev | pmid:22279377 |
| Davey MS et al. | Human neutrophil clearance of bacterial pathogens triggers anti-microbial γδ T cell responses in early infection. | 2011 | PLoS Pathog. | pmid:21589907 |
| Baumeister S et al. | Fosmidomycin uptake into Plasmodium and Babesia-infected erythrocytes is facilitated by parasite-induced new permeability pathways. | 2011 | PLoS ONE | pmid:21573242 |
| Meier S et al. | A transcriptional analysis of carotenoid, chlorophyll and plastidial isoprenoid biosynthesis genes during development and osmotic stress responses in Arabidopsis thaliana. | 2011 | BMC Syst Biol | pmid:21595952 |
| Lemuth K et al. | Engineering of a plasmid-free Escherichia coli strain for improved in vivo biosynthesis of astaxanthin. | 2011 | Microb. Cell Fact. | pmid:21521516 |
| Zhou K et al. | Novel reference genes for quantifying transcriptional responses of Escherichia coli to protein overexpression by quantitative PCR. | 2011 | BMC Mol. Biol. | pmid:21513543 |