| MeSH term | MeSH ID | Detail |
|---|---|---|
| Melanoma | D008545 | 69 associated lipids |
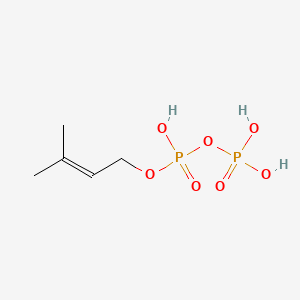
Dimethylallyl pyrophosphate
Dimethylallyl pyrophosphate is a lipid of Prenol Lipids (PR) class. Dimethylallyl pyrophosphate is associated with abnormalities such as Consumption-archaic term for TB and Wiskott-Aldrich Syndrome. The involved functions are known as Anabolism, Biochemical Pathway, Oxidation, Process and Chelating Activity [MoA]. Dimethylallyl pyrophosphate often locates in Chloroplasts, Plastids, chloroplast stroma, Cytosol and Cell membrane. The associated genes with Dimethylallyl pyrophosphate are IRF6 wt Allele and ADRBK1 gene. The related lipids are Sterols.
Cross Reference
Introduction
To understand associated biological information of Dimethylallyl pyrophosphate, we collected biological information of abnormalities, associated pathways, cellular/molecular locations, biological functions, related genes/proteins, lipids and common seen animal/experimental models with organized paragraphs from literatures.
What diseases are associated with Dimethylallyl pyrophosphate?
Dimethylallyl pyrophosphate is suspected in and other diseases in descending order of the highest number of associated sentences.
Related references are mostly published in these journals:
| Disease | Cross reference | Weighted score | Related literature |
|---|
Possible diseases from mapped MeSH terms on references
We collected disease MeSH terms mapped to the references associated with Dimethylallyl pyrophosphate
PubChem Associated disorders and diseases
What pathways are associated with Dimethylallyl pyrophosphate
Lipid pathways are not clear in current pathway databases. We organized associated pathways with Dimethylallyl pyrophosphate through full-text articles, including metabolic pathways or pathways of biological mechanisms.
Related references are published most in these journals:
| Pathway name | Related literatures |
|---|
PubChem Biomolecular Interactions and Pathways
Link to PubChem Biomolecular Interactions and PathwaysWhat cellular locations are associated with Dimethylallyl pyrophosphate?
Visualization in cellular structure
Associated locations are in red color. Not associated locations are in black.
Related references are published most in these journals:
| Location | Cross reference | Weighted score | Related literatures |
|---|
What functions are associated with Dimethylallyl pyrophosphate?
Related references are published most in these journals:
| Function | Cross reference | Weighted score | Related literatures |
|---|
What lipids are associated with Dimethylallyl pyrophosphate?
Related references are published most in these journals:
| Lipid concept | Cross reference | Weighted score | Related literatures |
|---|
What genes are associated with Dimethylallyl pyrophosphate?
Related references are published most in these journals:
| Gene | Cross reference | Weighted score | Related literatures |
|---|
What common seen animal models are associated with Dimethylallyl pyrophosphate?
There are no associated biomedical information in the current reference collection.
NCBI Entrez Crosslinks
All references with Dimethylallyl pyrophosphate
Download all related citations| Authors | Title | Published | Journal | PubMed Link |
|---|---|---|---|---|
| Liu T and Khosla C | Chemistry. A balancing act for Taxol precursor pathways in E. coli. | 2010 | Science | pmid:20929799 |
| Kiran U et al. | Structural and functional characterization of HMG-COA reductase from Artemisia annua. | 2010 | Bioinformation | pmid:21364776 |
| Dunne MR et al. | Preferential Th1 cytokine profile of phosphoantigen-stimulated human Vγ9Vδ2 T cells. | 2010 | Mediators Inflamm. | pmid:21403900 |
| George GM et al. | Virus-induced gene silencing of plastidial soluble inorganic pyrophosphatase impairs essential leaf anabolic pathways and reduces drought stress tolerance in Nicotiana benthamiana. | 2010 | Plant Physiol. | pmid:20605913 |
| Goto T et al. | Various Terpenoids Derived from Herbal and Dietary Plants Function as PPAR Modulators and Regulate Carbohydrate and Lipid Metabolism. | 2010 | PPAR Res | pmid:20613991 |
| Chu HM et al. | Binding and catalysis of Humulus lupulus adenylate isopentenyltransferase for the synthesis of isopentenylated diadenosine polyphosphates. | 2010 | FEBS Lett. | pmid:20807533 |
| Costa GG et al. | Transcriptome analysis of the oil-rich seed of the bioenergy crop Jatropha curcas L. | 2010 | BMC Genomics | pmid:20691070 |
| Thibodeaux CJ et al. | Linear free energy relationships demonstrate a catalytic role for the flavin mononucleotide coenzyme of the type II isopentenyl diphosphate:dimethylallyl diphosphate isomerase. | 2010 | J. Am. Chem. Soc. | pmid:20593767 |
| Andrews M et al. | The CaaX specificities of Arabidopsis protein prenyltransferases explain era1 and ggb phenotypes. | 2010 | BMC Plant Biol. | pmid:20565889 |
| Dellomonaco C et al. | The path to next generation biofuels: successes and challenges in the era of synthetic biology. | 2010 | Microb. Cell Fact. | pmid:20089184 |