| MeSH term | MeSH ID | Detail |
|---|---|---|
| Melanoma | D008545 | 69 associated lipids |
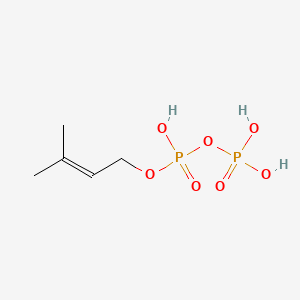
Dimethylallyl pyrophosphate
Dimethylallyl pyrophosphate is a lipid of Prenol Lipids (PR) class. Dimethylallyl pyrophosphate is associated with abnormalities such as Consumption-archaic term for TB and Wiskott-Aldrich Syndrome. The involved functions are known as Anabolism, Biochemical Pathway, Oxidation, Process and Chelating Activity [MoA]. Dimethylallyl pyrophosphate often locates in Chloroplasts, Plastids, chloroplast stroma, Cytosol and Cell membrane. The associated genes with Dimethylallyl pyrophosphate are IRF6 wt Allele and ADRBK1 gene. The related lipids are Sterols.
Cross Reference
Introduction
To understand associated biological information of Dimethylallyl pyrophosphate, we collected biological information of abnormalities, associated pathways, cellular/molecular locations, biological functions, related genes/proteins, lipids and common seen animal/experimental models with organized paragraphs from literatures.
What diseases are associated with Dimethylallyl pyrophosphate?
Dimethylallyl pyrophosphate is suspected in and other diseases in descending order of the highest number of associated sentences.
Related references are mostly published in these journals:
| Disease | Cross reference | Weighted score | Related literature |
|---|
Possible diseases from mapped MeSH terms on references
We collected disease MeSH terms mapped to the references associated with Dimethylallyl pyrophosphate
PubChem Associated disorders and diseases
What pathways are associated with Dimethylallyl pyrophosphate
Lipid pathways are not clear in current pathway databases. We organized associated pathways with Dimethylallyl pyrophosphate through full-text articles, including metabolic pathways or pathways of biological mechanisms.
Related references are published most in these journals:
| Pathway name | Related literatures |
|---|
PubChem Biomolecular Interactions and Pathways
Link to PubChem Biomolecular Interactions and PathwaysWhat cellular locations are associated with Dimethylallyl pyrophosphate?
Visualization in cellular structure
Associated locations are in red color. Not associated locations are in black.
Related references are published most in these journals:
| Location | Cross reference | Weighted score | Related literatures |
|---|
What functions are associated with Dimethylallyl pyrophosphate?
Related references are published most in these journals:
| Function | Cross reference | Weighted score | Related literatures |
|---|
What lipids are associated with Dimethylallyl pyrophosphate?
Related references are published most in these journals:
| Lipid concept | Cross reference | Weighted score | Related literatures |
|---|
What genes are associated with Dimethylallyl pyrophosphate?
Related references are published most in these journals:
| Gene | Cross reference | Weighted score | Related literatures |
|---|
What common seen animal models are associated with Dimethylallyl pyrophosphate?
There are no associated biomedical information in the current reference collection.
NCBI Entrez Crosslinks
All references with Dimethylallyl pyrophosphate
Download all related citations| Authors | Title | Published | Journal | PubMed Link |
|---|---|---|---|---|
| Brilli F et al. | Detection of plant volatiles after leaf wounding and darkening by proton transfer reaction "time-of-flight" mass spectrometry (PTR-TOF). | 2011 | PLoS ONE | pmid:21637822 |
| Mann FM et al. | Rv0989c encodes a novel (E)-geranyl diphosphate synthase facilitating decaprenyl diphosphate biosynthesis in Mycobacterium tuberculosis. | 2011 | FEBS Lett. | pmid:21237161 |
| Li Z et al. | Effect of temperature on postillumination isoprene emission in oak and poplar. | 2011 | Plant Physiol. | pmid:21177471 |
| Barrero RA et al. | De novo assembly of Euphorbia fischeriana root transcriptome identifies prostratin pathway related genes. | 2011 | BMC Genomics | pmid:22151917 |
| Tsang A et al. | Francisella tularensis 2-C-methyl-D-erythritol 4-phosphate cytidylyltransferase: kinetic characterization and phosphoregulation. | 2011 | PLoS ONE | pmid:21694781 |
| Misawa N | Carotenoid β-ring hydroxylase and ketolase from marine bacteria-promiscuous enzymes for synthesizing functional xanthophylls. | 2011 | Mar Drugs | pmid:21673887 |
| Bonitz T et al. | Evolutionary relationships of microbial aromatic prenyltransferases. | 2011 | PLoS ONE | pmid:22140437 |
| Clastre M et al. | Subcellular evidence for the involvement of peroxisomes in plant isoprenoid biosynthesis. | 2011 | Plant Signal Behav | pmid:22080790 |
| Hao da C et al. | The first insight into the tissue specific taxus transcriptome via Illumina second generation sequencing. | 2011 | PLoS ONE | pmid:21731678 |
| Tang Q et al. | An efficient approach to finding Siraitia grosvenorii triterpene biosynthetic genes by RNA-seq and digital gene expression analysis. | 2011 | BMC Genomics | pmid:21729270 |