| MeSH term | MeSH ID | Detail |
|---|---|---|
| Melanoma | D008545 | 69 associated lipids |
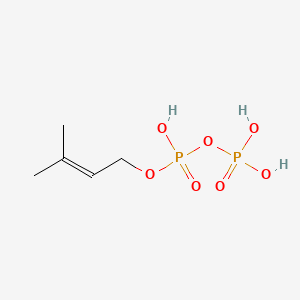
Dimethylallyl pyrophosphate
Dimethylallyl pyrophosphate is a lipid of Prenol Lipids (PR) class. Dimethylallyl pyrophosphate is associated with abnormalities such as Consumption-archaic term for TB and Wiskott-Aldrich Syndrome. The involved functions are known as Anabolism, Biochemical Pathway, Oxidation, Process and Chelating Activity [MoA]. Dimethylallyl pyrophosphate often locates in Chloroplasts, Plastids, chloroplast stroma, Cytosol and Cell membrane. The associated genes with Dimethylallyl pyrophosphate are IRF6 wt Allele and ADRBK1 gene. The related lipids are Sterols.
Cross Reference
Introduction
To understand associated biological information of Dimethylallyl pyrophosphate, we collected biological information of abnormalities, associated pathways, cellular/molecular locations, biological functions, related genes/proteins, lipids and common seen animal/experimental models with organized paragraphs from literatures.
What diseases are associated with Dimethylallyl pyrophosphate?
Dimethylallyl pyrophosphate is suspected in and other diseases in descending order of the highest number of associated sentences.
Related references are mostly published in these journals:
| Disease | Cross reference | Weighted score | Related literature |
|---|
Possible diseases from mapped MeSH terms on references
We collected disease MeSH terms mapped to the references associated with Dimethylallyl pyrophosphate
PubChem Associated disorders and diseases
What pathways are associated with Dimethylallyl pyrophosphate
Lipid pathways are not clear in current pathway databases. We organized associated pathways with Dimethylallyl pyrophosphate through full-text articles, including metabolic pathways or pathways of biological mechanisms.
Related references are published most in these journals:
| Pathway name | Related literatures |
|---|
PubChem Biomolecular Interactions and Pathways
Link to PubChem Biomolecular Interactions and PathwaysWhat cellular locations are associated with Dimethylallyl pyrophosphate?
Visualization in cellular structure
Associated locations are in red color. Not associated locations are in black.
Related references are published most in these journals:
| Location | Cross reference | Weighted score | Related literatures |
|---|
What functions are associated with Dimethylallyl pyrophosphate?
Related references are published most in these journals:
| Function | Cross reference | Weighted score | Related literatures |
|---|
What lipids are associated with Dimethylallyl pyrophosphate?
Related references are published most in these journals:
| Lipid concept | Cross reference | Weighted score | Related literatures |
|---|
What genes are associated with Dimethylallyl pyrophosphate?
Related references are published most in these journals:
| Gene | Cross reference | Weighted score | Related literatures |
|---|
What common seen animal models are associated with Dimethylallyl pyrophosphate?
There are no associated biomedical information in the current reference collection.
NCBI Entrez Crosslinks
All references with Dimethylallyl pyrophosphate
Download all related citations| Authors | Title | Published | Journal | PubMed Link |
|---|---|---|---|---|
| pmid:24692239 | ||||
| Ghirardo A et al. | Metabolic flux analysis of plastidic isoprenoid biosynthesis in poplar leaves emitting and nonemitting isoprene. | 2014 | Plant Physiol. | pmid:24590857 |
| pmid:24430210 | ||||
| Chen X et al. | Statistical experimental design guided optimization of a one-pot biphasic multienzyme total synthesis of amorpha-4,11-diene. | 2013 | PLoS ONE | pmid:24278153 |
| Rudolf JD and Poulter CD | Tyrosine O-prenyltransferase SirD catalyzes S-, C-, and N-prenylations on tyrosine and tryptophan derivatives. | 2013 | ACS Chem. Biol. | pmid:24083562 |
| Rasulov B et al. | Competition between isoprene emission and pigment synthesis during leaf development in aspen. | 2014 | Plant Cell Environ. | pmid:24033429 |
| Chang WC et al. | Current development in isoprenoid precursor biosynthesis and regulation. | 2013 | Curr Opin Chem Biol | pmid:23891475 |
| Zhou C et al. | Isopentenyl diphosphate and dimethylallyl diphosphate/isopentenyl diphosphate ratio measured with recombinant isopentenyl diphosphate isomerase and isoprene synthase. | 2013 | Anal. Biochem. | pmid:23747531 |
| Zhao L et al. | Methylerythritol phosphate pathway of isoprenoid biosynthesis. | 2013 | Annu. Rev. Biochem. | pmid:23746261 |
| Banerjee A et al. | Feedback inhibition of deoxy-D-xylulose-5-phosphate synthase regulates the methylerythritol 4-phosphate pathway. | 2013 | J. Biol. Chem. | pmid:23612965 |