| MeSH term | MeSH ID | Detail |
|---|---|---|
| Melanoma | D008545 | 69 associated lipids |
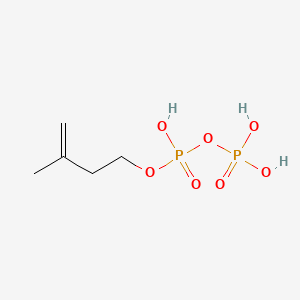
Isopentenyl pyrophosphate
Isopentenyl pyrophosphate is a lipid of Prenol Lipids (PR) class. Isopentenyl pyrophosphate is associated with abnormalities such as Tuberculosis, NAVAJO NEUROHEPATOPATHY and Cryptosporidiosis. The involved functions are known as Signal, Anabolism, T-Cell Activation, T-Cell Proliferation and isoprenoid biosynthetic process. Isopentenyl pyrophosphate often locates in Protoplasm, Host Cell, soluble, Plastids and Cell surface. The associated genes with Isopentenyl pyrophosphate are oxytocin, 1-desamino-(O-Et-Tyr)(2)-, HLA-E gene, RAP1A gene, Gene Family and Orthologous Gene. The related lipids are Steroids, Sterols and isopentenol.
Cross Reference
Introduction
To understand associated biological information of Isopentenyl pyrophosphate, we collected biological information of abnormalities, associated pathways, cellular/molecular locations, biological functions, related genes/proteins, lipids and common seen animal/experimental models with organized paragraphs from literatures.
What diseases are associated with Isopentenyl pyrophosphate?
Isopentenyl pyrophosphate is suspected in Tuberculosis, NAVAJO NEUROHEPATOPATHY, Cryptosporidiosis and other diseases in descending order of the highest number of associated sentences.
Related references are mostly published in these journals:
| Disease | Cross reference | Weighted score | Related literature |
|---|
Possible diseases from mapped MeSH terms on references
We collected disease MeSH terms mapped to the references associated with Isopentenyl pyrophosphate
PubChem Associated disorders and diseases
What pathways are associated with Isopentenyl pyrophosphate
Lipid pathways are not clear in current pathway databases. We organized associated pathways with Isopentenyl pyrophosphate through full-text articles, including metabolic pathways or pathways of biological mechanisms.
Related references are published most in these journals:
| Pathway name | Related literatures |
|---|
PubChem Biomolecular Interactions and Pathways
Link to PubChem Biomolecular Interactions and PathwaysWhat cellular locations are associated with Isopentenyl pyrophosphate?
Visualization in cellular structure
Associated locations are in red color. Not associated locations are in black.
Related references are published most in these journals:
| Location | Cross reference | Weighted score | Related literatures |
|---|
What functions are associated with Isopentenyl pyrophosphate?
Related references are published most in these journals:
| Function | Cross reference | Weighted score | Related literatures |
|---|
What lipids are associated with Isopentenyl pyrophosphate?
Related references are published most in these journals:
| Lipid concept | Cross reference | Weighted score | Related literatures |
|---|
What genes are associated with Isopentenyl pyrophosphate?
Related references are published most in these journals:
| Gene | Cross reference | Weighted score | Related literatures |
|---|
What common seen animal models are associated with Isopentenyl pyrophosphate?
There are no associated biomedical information in the current reference collection.
NCBI Entrez Crosslinks
All references with Isopentenyl pyrophosphate
Download all related citations| Authors | Title | Published | Journal | PubMed Link |
|---|---|---|---|---|
| Schilmiller AL et al. | Monoterpenes in the glandular trichomes of tomato are synthesized from a neryl diphosphate precursor rather than geranyl diphosphate. | 2009 | Proc. Natl. Acad. Sci. U.S.A. | pmid:19487664 |
| Liu YL et al. | Taxodione and arenarone inhibit farnesyl diphosphate synthase by binding to the isopentenyl diphosphate site. | 2014 | Proc. Natl. Acad. Sci. U.S.A. | pmid:24927548 |
| Bohlmann J and Gershenzon J | Old substrates for new enzymes of terpenoid biosynthesis. | 2009 | Proc. Natl. Acad. Sci. U.S.A. | pmid:19553206 |
| Leonard E et al. | Combining metabolic and protein engineering of a terpenoid biosynthetic pathway for overproduction and selectivity control. | 2010 | Proc. Natl. Acad. Sci. U.S.A. | pmid:20643967 |
| Lange BM et al. | Isoprenoid biosynthesis: the evolution of two ancient and distinct pathways across genomes. | 2000 | Proc. Natl. Acad. Sci. U.S.A. | pmid:11078528 |
| Henry LK et al. | Orthologs of the archaeal isopentenyl phosphate kinase regulate terpenoid production in plants. | 2015 | Proc. Natl. Acad. Sci. U.S.A. | pmid:26216978 |
| Dudareva N et al. | The nonmevalonate pathway supports both monoterpene and sesquiterpene formation in snapdragon flowers. | 2005 | Proc. Natl. Acad. Sci. U.S.A. | pmid:15630092 |
| Sun Z et al. | Differential expression of two isopentenyl pyrophosphate isomerases and enhanced carotenoid accumulation in a unicellular chlorophyte. | 1998 | Proc. Natl. Acad. Sci. U.S.A. | pmid:9736763 |
| Takahashi S et al. | A 1-deoxy-D-xylulose 5-phosphate reductoisomerase catalyzing the formation of 2-C-methyl-D-erythritol 4-phosphate in an alternative nonmevalonate pathway for terpenoid biosynthesis. | 1998 | Proc. Natl. Acad. Sci. U.S.A. | pmid:9707569 |
| Catusse J et al. | Proteome-wide characterization of sugarbeet seed vigor and its tissue specific expression. | 2008 | Proc. Natl. Acad. Sci. U.S.A. | pmid:18635686 |