| Authors | Title | Published | Journal | PubMed Link |
|---|---|---|---|---|
| pmid:13088191 | ||||
| pmid:13492088 | ||||
| pmid:25712566 | ||||
| pmid:25655771 | ||||
| pmid:25584873 | ||||
| pmid:25564760 | ||||
| pmid:20273114 | ||||
| pmid:15403928 | ||||
| pmid:17833646 | ||||
| pmid:25721651 |
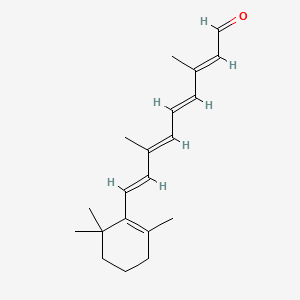
Retinal
Retinal is a lipid of Prenol Lipids (PR) class. Retinal is associated with abnormalities such as Retinal Detachment, Uveitis, Endophthalmitis, Infection and LEBER CONGENITAL AMAUROSIS, TYPE II (disorder). The involved functions are known as Pigment, immunoreactivity, Infiltration, Energy Absorption and chromophore. Retinal often locates in Entire visual system, Autosome, Membrane, Basement membrane and Thoracic region (surface region of back). The associated genes with Retinal are isorhodopsin, RPE65 gene, RND1 gene, RPE gene and CASP8AP2 gene. The related lipids are Phosphatidylserines, Membrane Lipids, Fatty Acids, Liposomes and oxidized lipid. The related experimental models are Mouse Model, Knock-out and Genetically Engineered Mouse.
Cross Reference
Introduction
To understand associated biological information of Retinal, we collected biological information of abnormalities, associated pathways, cellular/molecular locations, biological functions, related genes/proteins, lipids and common seen animal/experimental models with organized paragraphs from literatures.
What diseases are associated with Retinal?
Retinal is suspected in Retinal Degeneration, Retinal Diseases, Retinal Dystrophies, Leber Congenital Amaurosis, Retinitis Pigmentosa, Stargardt's disease and other diseases in descending order of the highest number of associated sentences.
Related references are mostly published in these journals:
- J. Biol. Chem. (57)
- Invest. Ophthalmol. Vis. Sci. (28)
- Proc. Natl. Acad. Sci. U.S.A. (22)
- Others (140)
| Disease | Cross reference | Weighted score | Related literature |
|---|
No disease MeSH terms mapped to the current reference collection.
PubChem Associated disorders and diseases
What pathways are associated with Retinal
There are no associated biomedical information in the current reference collection.
PubChem Biomolecular Interactions and Pathways
Link to PubChem Biomolecular Interactions and PathwaysWhat cellular locations are associated with Retinal?
Visualization in cellular structure
Associated locations are in red color. Not associated locations are in black.
Related references are published most in these journals:
- J. Biol. Chem. (52)
- Proc. Natl. Acad. Sci. U.S.A. (22)
- Invest. Ophthalmol. Vis. Sci. (20)
- Others (136)
| Location | Cross reference | Weighted score | Related literatures |
|---|
What functions are associated with Retinal?
Related references are published most in these journals:
| Function | Cross reference | Weighted score | Related literatures |
|---|
What lipids are associated with Retinal?
Related references are published most in these journals:
| Lipid concept | Cross reference | Weighted score | Related literatures |
|---|
What genes are associated with Retinal?
Related references are published most in these journals:
- J. Biol. Chem. (63)
- Invest. Ophthalmol. Vis. Sci. (21)
- Proc. Natl. Acad. Sci. U.S.A. (20)
- Others (145)
| Gene | Cross reference | Weighted score | Related literatures |
|---|
What common seen animal models are associated with Retinal?
Mouse Model
Mouse Model are used in the study 'Improvement in rod and cone function in mouse model of Fundus albipunctatus after pharmacologic treatment with 9-cis-retinal.' (Maeda A et al., 2006), Mouse Model are used in the study 'Quantitative mapping of ion channel regulation by visual cycle activity in rodent photoreceptors in vivo.' (Berkowitz BA et al., 2009), Mouse Model are used in the study 'Chemistry and biology of vision.' (Palczewski K, 2012) and Mouse Model are used in the study 'Photoreceptor proteins initiate microglial activation via Toll-like receptor 4 in retinal degeneration mediated by all-trans-retinal.' (Kohno H et al., 2013).
Knock-out
Knock-out are used in the study 'Conditional Ablation of Retinol Dehydrogenase 10 in the Retinal Pigmented Epithelium Causes Delayed Dark Adaption in Mice.' (Sahu B et al., 2015), Knock-out are used in the study 'Interpretations of fundus autofluorescence from studies of the bisretinoids of the retina.' (Sparrow JR et al., 2010), Knock-out are used in the study 'Bacterioopsin-mediated regulation of bacterioruberin biosynthesis in Halobacterium salinarum.' (Dummer AM et al., 2011) and Knock-out are used in the study 'RPE65 is essential for the function of cone photoreceptors in NRL-deficient mice.' (Wenzel A et al., 2007).
Genetically Engineered Mouse
Genetically Engineered Mouse are used in the study 'Limited roles of Rdh8, Rdh12, and Abca4 in all-trans-retinal clearance in mouse retina.' (Maeda A et al., 2009) and Genetically Engineered Mouse are used in the study 'Recovery of visual functions in a mouse model of Leber congenital amaurosis.' (Van Hooser JP et al., 2002).
Related references are published most in these journals:
| Model | Cross reference | Weighted score | Related literatures |
|---|