| MeSH term | MeSH ID | Detail |
|---|---|---|
| Limb Deformities, Congenital | D017880 | 4 associated lipids |
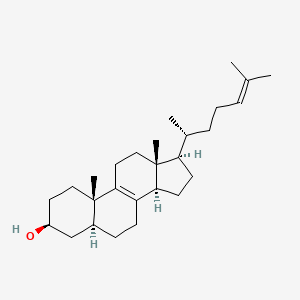
Zymosterol
Zymosterol is a lipid of Sterol Lipids (ST) class. Zymosterol is associated with abnormalities such as Congenital Abnormality, Coronary Arteriosclerosis, Liver diseases, Alzheimer's Disease and Virus Diseases. The involved functions are known as Process, enzyme activity, Anabolism, Transcription, Genetic and Synthesis. Zymosterol often locates in Cytosol, Mitochondria, Articular system, Protoplasm and Tissue membrane. The associated genes with Zymosterol are Genome, Homologous Gene, Amino Acids, Acidic, Peptide Fragments and DLC1 gene. The related lipids are zymosterol, Cardiolipins, 25-azacholesterol, Sterols and Steroids. The related experimental models are Knock-out.
Cross Reference
Introduction
To understand associated biological information of Zymosterol, we collected biological information of abnormalities, associated pathways, cellular/molecular locations, biological functions, related genes/proteins, lipids and common seen animal/experimental models with organized paragraphs from literatures.
What diseases are associated with Zymosterol?
Zymosterol is suspected in Leishmaniasis, Congenital Abnormality, Coronary Arteriosclerosis, Liver diseases, Alzheimer's Disease, Virus Diseases and other diseases in descending order of the highest number of associated sentences.
Related references are mostly published in these journals:
| Disease | Cross reference | Weighted score | Related literature |
|---|
Possible diseases from mapped MeSH terms on references
We collected disease MeSH terms mapped to the references associated with Zymosterol
PubChem Associated disorders and diseases
What pathways are associated with Zymosterol
There are no associated biomedical information in the current reference collection.
PubChem Biomolecular Interactions and Pathways
Link to PubChem Biomolecular Interactions and PathwaysWhat cellular locations are associated with Zymosterol?
Visualization in cellular structure
Associated locations are in red color. Not associated locations are in black.
Related references are published most in these journals:
| Location | Cross reference | Weighted score | Related literatures |
|---|
What functions are associated with Zymosterol?
Related references are published most in these journals:
| Function | Cross reference | Weighted score | Related literatures |
|---|
What lipids are associated with Zymosterol?
Related references are published most in these journals:
| Lipid concept | Cross reference | Weighted score | Related literatures |
|---|
What genes are associated with Zymosterol?
Related references are published most in these journals:
| Gene | Cross reference | Weighted score | Related literatures |
|---|
What common seen animal models are associated with Zymosterol?
Knock-out
Knock-out are used in the study 'Systems analysis of metabolism in the pathogenic trypanosomatid Leishmania major.' (Chavali AK et al., 2008).
Related references are published most in these journals:
| Model | Cross reference | Weighted score | Related literatures |
|---|
NCBI Entrez Crosslinks
All references with Zymosterol
Download all related citations| Authors | Title | Published | Journal | PubMed Link |
|---|---|---|---|---|
| Iwaki T et al. | Multiple functions of ergosterol in the fission yeast Schizosaccharomyces pombe. | 2008 | Microbiology (Reading, Engl.) | pmid:18310029 |
| Urbina JA et al. | Modification of the sterol composition of Trypanosoma (Schizotrypanum) cruzi epimastigotes by delta 24(25)-sterol methyl transferase inhibitors and their combinations with ketoconazole. | 1995 | Mol. Biochem. Parasitol. | pmid:8577328 |
| Steffensen KR et al. | Reduced fertility and inability of oocytes to resume meiosis in mice deficient of the Lxr genes. | 2006 | Mol. Cell. Endocrinol. | pmid:16895745 |
| Marijanovic Z et al. | Closing the gap: identification of human 3-ketosteroid reductase, the last unknown enzyme of mammalian cholesterol biosynthesis. | 2003 | Mol. Endocrinol. | pmid:12829805 |
| Liu Y et al. | Cytochrome P450 17alpha hydroxylase/17,20 lyase (CYP17) function in cholesterol biosynthesis: identification of squalene monooxygenase (epoxidase) activity associated with CYP17 in Leydig cells. | 2005 | Mol. Endocrinol. | pmid:15761033 |
| Kaneshiro ES et al. | The Pneumocystis carinii drug target S-adenosyl-L-methionine:sterol C-24 methyl transferase has a unique substrate preference. | 2002 | Mol. Microbiol. | pmid:12010494 |
| Andersen MR et al. | Metabolic model integration of the bibliome, genome, metabolome and reactome of Aspergillus niger. | 2008 | Mol. Syst. Biol. | pmid:18364712 |
| Chavali AK et al. | Systems analysis of metabolism in the pathogenic trypanosomatid Leishmania major. | 2008 | Mol. Syst. Biol. | pmid:18364711 |
| Blanc M et al. | Host defense against viral infection involves interferon mediated down-regulation of sterol biosynthesis. | 2011 | PLoS Biol. | pmid:21408089 |
| Almaas E et al. | The activity reaction core and plasticity of metabolic networks. | 2005 | PLoS Comput. Biol. | pmid:16362071 |