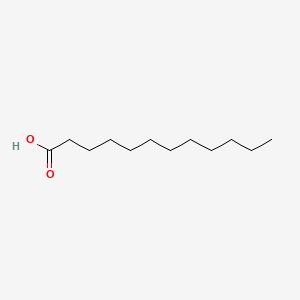
lauric acid
lauric acid is a lipid of Fatty Acyls (FA) class. Lauric acid is associated with abnormalities such as Infection, Renal tubular disorder, Hypertensive disease, Obesity and Mycoses. The involved functions are known as Transcription, Genetic, Signal Transduction, Mutation, metaplastic cell transformation and Anabolism. Lauric acid often locates in Skin, Plasma membrane, Cytoplasmic matrix, Body tissue and Palmar surface. The associated genes with lauric acid are Gene Family, SLC33A1 gene, Homologous Gene, Open Reading Frames and P4HTM gene. The related lipids are Fatty Acids, Oleic Acids, Palmitates, Stearates and 9,11-linoleic acid.
Cross Reference
Introduction
To understand associated biological information of lauric acid, we collected biological information of abnormalities, associated pathways, cellular/molecular locations, biological functions, related genes/proteins, lipids and common seen animal/experimental models with organized paragraphs from literatures.
What diseases are associated with lauric acid?
lauric acid is suspected in Renal tubular disorder, Hypertensive disease, Infection, Renal vascular disorder, Obesity, Mycoses and other diseases in descending order of the highest number of associated sentences.
Related references are mostly published in these journals:
| Disease | Cross reference | Weighted score | Related literature |
|---|
Possible diseases from mapped MeSH terms on references
We collected disease MeSH terms mapped to the references associated with lauric acid
PubChem Associated disorders and diseases
What pathways are associated with lauric acid
There are no associated biomedical information in the current reference collection.
PubChem Biomolecular Interactions and Pathways
Link to PubChem Biomolecular Interactions and PathwaysWhat cellular locations are associated with lauric acid?
Visualization in cellular structure
Associated locations are in red color. Not associated locations are in black.
Related references are published most in these journals:
| Location | Cross reference | Weighted score | Related literatures |
|---|
What functions are associated with lauric acid?
Related references are published most in these journals:
| Function | Cross reference | Weighted score | Related literatures |
|---|
What lipids are associated with lauric acid?
Related references are published most in these journals:
| Lipid concept | Cross reference | Weighted score | Related literatures |
|---|
What genes are associated with lauric acid?
Related references are published most in these journals:
| Gene | Cross reference | Weighted score | Related literatures |
|---|
What common seen animal models are associated with lauric acid?
There are no associated biomedical information in the current reference collection.
NCBI Entrez Crosslinks
All references with lauric acid
Download all related citations| Authors | Title | Published | Journal | PubMed Link |
|---|---|---|---|---|
| Gawas SD et al. | Solvent Free Lipase Catalysed Synthesis of Ethyl Laurate: Optimization and Kinetic Studies. | 2016 | Appl. Biochem. Biotechnol. | pmid:27470112 |
| Takenaka S et al. | Adaptation of Pseudomonas sp. strain 7-6 to quaternary ammonium compounds and their degradation via dual pathways. | 2007 | Appl. Environ. Microbiol. | pmid:17261523 |
| Schallmey A et al. | Characterization of cytochrome P450 monooxygenase CYP154H1 from the thermophilic soil bacterium Thermobifida fusca. | 2011 | Appl. Microbiol. Biotechnol. | pmid:21057946 |
| Beyer N et al. | P450 fused to phosphite dehydrogenase allows phosphite-driven selective oxidations. | 2017 | Appl. Microbiol. Biotechnol. | pmid:27900443 |
| Ren Q et al. | Enatiomerically pure hydroxycarboxylic acids: current approaches and future perspectives. | 2010 | Appl. Microbiol. Biotechnol. | pmid:20393709 |
| Hama S et al. | Transesterification of phosphatidylcholine in sn-1 position through direct use of lipase-producing Rhizopus oryzae cells as whole-cell biocatalyst. | 2011 | Appl. Microbiol. Biotechnol. | pmid:21468705 |
| Lu XY et al. | Production of poly(3-hydroxybutyrate- co-3-hydroxyhexanoate) with flexible 3-hydroxyhexanoate content in Aeromonas hydrophila CGMCC 0911. | 2004 | Appl. Microbiol. Biotechnol. | pmid:12920488 |
| Chung A et al. | Microbial production of 3-hydroxydodecanoic acid by pha operon and fadBA knockout mutant of Pseudomonas putida KT2442 harboring tesB gene. | 2009 | Appl. Microbiol. Biotechnol. | pmid:19271216 |
| Chen GQ et al. | Industrial scale production of poly(3-hydroxybutyrate-co-3-hydroxyhexanoate). | 2001 | Appl. Microbiol. Biotechnol. | pmid:11693933 |
| Kojima K et al. | A simple method for isolation and construction of markerless cyanobacterial mutants defective in acyl-acyl carrier protein synthetase. | 2016 | Appl. Microbiol. Biotechnol. | pmid:27704180 |
| Sato S et al. | Expression and characterization of (R)-specific enoyl coenzyme A hydratases making a channeling route to polyhydroxyalkanoate biosynthesis in Pseudomonas putida. | 2011 | Appl. Microbiol. Biotechnol. | pmid:21327961 |
| Di Bello D et al. | Presence and inducibility by beta-naphthoflavone of CYP1A1, CYP1B1 and phase II enzymes in Trematomus bernacchii, an Antarctic fish. | 2007 | Aquat. Toxicol. | pmid:17643506 |
| Nabb DL et al. | Comparison of basal level metabolic enzyme activities of freshly isolated hepatocytes from rainbow trout (Oncorhynchus mykiss) and rat. | 2006 | Aquat. Toxicol. | pmid:16935359 |
| Thibaut R et al. | Regio-specific hydroxylation of nonylphenol and the involvement of CYP2K- and CYP2M-like iso-enzymes in Atlantic salmon (Salmo salar). | 2002 | Aquat. Toxicol. | pmid:11792434 |
| Dohme F et al. | Digestive and metabolic utilization of lauric, myristic and stearic acid in cows, and associated effects on milk fat quality. | 2004 | Arch Anim Nutr | pmid:15195905 |
| Geiger F et al. | Trochanteric fractures in the elderly: the influence of primary hip arthroplasty on 1-year mortality. | 2007 | Arch Orthop Trauma Surg | pmid:17899138 |
| Berschauer F et al. | [Nutritive physiological effect of dietary fats in growing swine rations. 3. Effect of sunflower and coconut kernels on protein and fat retention, backfat fatty acid pattern and various blood parameters in piglets]. | 1983 | Arch Tierernahr | pmid:6370196 |
| Helali E and Hashem H | [Qualitative and quantitative studies on the metabolism of fatty acids, in particular on that of linoleic acid in calves]. | 1975 | Arch Tierernahr | pmid:1233966 |
| Kolattukudy PE and Rogers L | Biosynthesis of 3-hydroxy fatty acids, the pheromone components of female mallard ducks, by cell-free preparations from the uropygial gland. | 1987 | Arch. Biochem. Biophys. | pmid:3813530 |
| Lewis DF | Essential requirements for substrate binding affinity and selectivity toward human CYP2 family enzymes. | 2003 | Arch. Biochem. Biophys. | pmid:12464242 |