| MeSH term | MeSH ID | Detail |
|---|---|---|
| Atherosclerosis | D050197 | 85 associated lipids |
| Vascular System Injuries | D057772 | 2 associated lipids |
| Peripheral Arterial Disease | D058729 | 7 associated lipids |
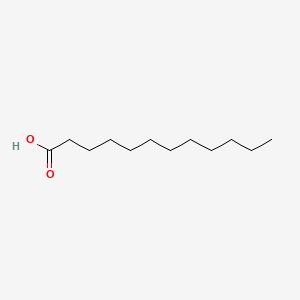
lauric acid
lauric acid is a lipid of Fatty Acyls (FA) class. Lauric acid is associated with abnormalities such as Infection, Renal tubular disorder, Hypertensive disease, Obesity and Mycoses. The involved functions are known as Transcription, Genetic, Signal Transduction, Mutation, metaplastic cell transformation and Anabolism. Lauric acid often locates in Skin, Plasma membrane, Cytoplasmic matrix, Body tissue and Palmar surface. The associated genes with lauric acid are Gene Family, SLC33A1 gene, Homologous Gene, Open Reading Frames and P4HTM gene. The related lipids are Fatty Acids, Oleic Acids, Palmitates, Stearates and 9,11-linoleic acid.
Cross Reference
Introduction
To understand associated biological information of lauric acid, we collected biological information of abnormalities, associated pathways, cellular/molecular locations, biological functions, related genes/proteins, lipids and common seen animal/experimental models with organized paragraphs from literatures.
What diseases are associated with lauric acid?
lauric acid is suspected in Renal tubular disorder, Hypertensive disease, Infection, Renal vascular disorder, Obesity, Mycoses and other diseases in descending order of the highest number of associated sentences.
Related references are mostly published in these journals:
| Disease | Cross reference | Weighted score | Related literature |
|---|
Possible diseases from mapped MeSH terms on references
We collected disease MeSH terms mapped to the references associated with lauric acid
PubChem Associated disorders and diseases
What pathways are associated with lauric acid
There are no associated biomedical information in the current reference collection.
PubChem Biomolecular Interactions and Pathways
Link to PubChem Biomolecular Interactions and PathwaysWhat cellular locations are associated with lauric acid?
Visualization in cellular structure
Associated locations are in red color. Not associated locations are in black.
Related references are published most in these journals:
| Location | Cross reference | Weighted score | Related literatures |
|---|
What functions are associated with lauric acid?
Related references are published most in these journals:
| Function | Cross reference | Weighted score | Related literatures |
|---|
What lipids are associated with lauric acid?
Related references are published most in these journals:
| Lipid concept | Cross reference | Weighted score | Related literatures |
|---|
What genes are associated with lauric acid?
Related references are published most in these journals:
| Gene | Cross reference | Weighted score | Related literatures |
|---|
What common seen animal models are associated with lauric acid?
There are no associated biomedical information in the current reference collection.
NCBI Entrez Crosslinks
All references with lauric acid
Download all related citations| Authors | Title | Published | Journal | PubMed Link |
|---|---|---|---|---|
| Rottensteiner H et al. | The peroxisomal transporter gene ANT1 is regulated by a deviant oleate response element (ORE): characterization of the signal for fatty acid induction. | 2002 | Biochem. J. | pmid:12071844 |
| Whiting P and Dinan L | Formation of apolar ecdysteroid conjugates by ovaries of the house cricket Acheta domesticus in vitro. | 1988 | Biochem. J. | pmid:3421912 |
| Girvan HM et al. | Novel haem co-ordination variants of flavocytochrome P450BM3. | 2009 | Biochem. J. | pmid:18721129 |
| Gibson GG et al. | Cytochrome P-450 induction by clofibrate. Purification and properties of a hepatic cytochrome P-450 relatively specific for the 12- and 11-hydroxylation of dodecanoic acid (lauric acid). | 1982 | Biochem. J. | pmid:7103935 |
| Hasegawa K et al. | Growth of beta(2)-microglobulin-related amyloid fibrils by non-esterified fatty acids at a neutral pH. | 2008 | Biochem. J. | pmid:18637792 |
| VAN DE KAMER JH et al. | Gas-liquid partition chromatography: the separation and micro-estimation of volatile fatty acids from formic acid to dodecanoic acid. | 1955 | Biochem. J. | pmid:13260193 |
| JAMES AT and MARTIN AJ | Gas-liquid partition chromatography; the separation and micro-estimation of volatile fatty acids from formic acid to dodecanoic acid. | 1952 | Biochem. J. | pmid:14934673 |
| Girvan HM et al. | Glutamate-haem ester bond formation is disfavoured in flavocytochrome P450 BM3: characterization of glutamate substitution mutants at the haem site of P450 BM3. | 2010 | Biochem. J. | pmid:20180779 |
| Lind KE and Moller JV | Ligand-apomyoglobin interactions. Configurational adaptability of the haem-binding site. | 1976 | Biochem. J. | pmid:949328 |
| Muller DN et al. | Mouse Cyp4a isoforms: enzymatic properties, gender- and strain-specific expression, and role in renal 20-hydroxyeicosatetraenoic acid formation. | 2007 | Biochem. J. | pmid:17112342 |