| MeSH term | MeSH ID | Detail |
|---|---|---|
| Bacterial Infections | D001424 | 21 associated lipids |
| Body Weight | D001835 | 333 associated lipids |
| Drug Hypersensitivity | D004342 | 20 associated lipids |
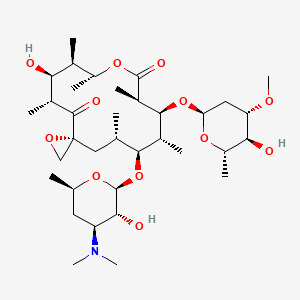
oleandomycin
oleandomycin is a lipid of Polyketides (PK) class. The involved functions are known as 5-(carboxyamino)imidazole ribonucleotide mutase activity, Methylation, enzyme activity, Anabolism and Biosynthetic Pathways. Oleandomycin often locates in Chromosomes and Protoplasm. The associated genes with oleandomycin are resistance genes, Open Reading Frames, CTTNBP2 gene, Gene Feature and Gene Clusters.
Cross Reference
Introduction
To understand associated biological information of oleandomycin, we collected biological information of abnormalities, associated pathways, cellular/molecular locations, biological functions, related genes/proteins, lipids and common seen animal/experimental models with organized paragraphs from literatures.
What diseases are associated with oleandomycin?
There are no associated biomedical information in the current reference collection.
Possible diseases from mapped MeSH terms on references
We collected disease MeSH terms mapped to the references associated with oleandomycin
PubChem Associated disorders and diseases
What pathways are associated with oleandomycin
There are no associated biomedical information in the current reference collection.
PubChem Biomolecular Interactions and Pathways
Link to PubChem Biomolecular Interactions and PathwaysWhat cellular locations are associated with oleandomycin?
Visualization in cellular structure
Associated locations are in red color. Not associated locations are in black.
Related references are published most in these journals:
| Location | Cross reference | Weighted score | Related literatures |
|---|
What functions are associated with oleandomycin?
Related references are published most in these journals:
| Function | Cross reference | Weighted score | Related literatures |
|---|
What lipids are associated with oleandomycin?
There are no associated biomedical information in the current reference collection.
What genes are associated with oleandomycin?
Related references are published most in these journals:
| Gene | Cross reference | Weighted score | Related literatures |
|---|
What common seen animal models are associated with oleandomycin?
There are no associated biomedical information in the current reference collection.
NCBI Entrez Crosslinks
All references with oleandomycin
Download all related citations| Authors | Title | Published | Journal | PubMed Link |
|---|---|---|---|---|
| Shevchenko AA et al. | [Biosynthesis of oleandomycin by cultures of Streptomyces antibioticus obtained after regeneration and fusion of protoplasts]. | 1989 Mar-Apr | Mikrobiol. Zh. | pmid:2668708 |
| Sartori E et al. | In vitro interaction of rat liver cytochromes P-450 with erythromycin, oleandomycin and erythralosamine derivatives. Importance of structural factors. | 1989 | Biochem. Pharmacol. | pmid:2735945 |
| Marshall VP et al. | Microbial O-phosphorylation of macrolide antibiotics. | 1989 | J. Antibiot. | pmid:2921218 |
| Labro MT et al. | Comparison of the in-vitro effect of several macrolides on the oxidative burst of human neutrophils. | 1989 | J. Antimicrob. Chemother. | pmid:2559072 |
| Tankevich ME et al. | [Mutagenic effect of mitomycin C on different strains of oleandomycin producer Streptomyces antibioticus]. | 1989 | Antibiot. Khimioter. | pmid:2517387 |
| Beliakova IuG et al. | [Oleandomycin and its natural and synthetic analogs. I. Dissociative ionization of oleandomycin and its derivatives]. | 1989 | Antibiot. Khimioter. | pmid:2751374 |
| Beliakova IuG et al. | [Oleandomycin and its natural and synthetic analogs. II. Structure and properties of oleandomycin B, a new compound of the oleandomycin group]. | 1989 | Antibiot. Khimioter. | pmid:2751375 |
| Serova IP et al. | [Study of the process of inactivating oleandomycin phosphate preparations]. | 1989 | Antibiot. Khimioter. | pmid:2751382 |
| Borisenko KK et al. | [The immediate results of treating patients with infectious forms of syphilis by the abbreviated single-course reserve method]. | 1989 | Vestn Dermatol Venerol | pmid:2816017 |
| Petrova TB and Savitskaia TN | [Prenatal effect of oleandomycin on the development of the immunogenesis organs]. | 1988 May-Jun | Farmakol Toksikol | pmid:3410038 |