| MeSH term | MeSH ID | Detail |
|---|---|---|
| Bacterial Infections | D001424 | 21 associated lipids |
| Body Weight | D001835 | 333 associated lipids |
| Drug Hypersensitivity | D004342 | 20 associated lipids |
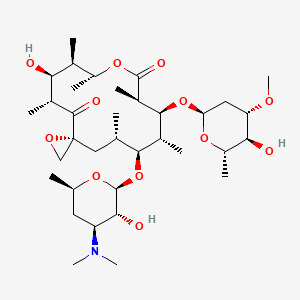
oleandomycin
oleandomycin is a lipid of Polyketides (PK) class. The involved functions are known as 5-(carboxyamino)imidazole ribonucleotide mutase activity, Methylation, enzyme activity, Anabolism and Biosynthetic Pathways. Oleandomycin often locates in Chromosomes and Protoplasm. The associated genes with oleandomycin are resistance genes, Open Reading Frames, CTTNBP2 gene, Gene Feature and Gene Clusters.
Cross Reference
Introduction
To understand associated biological information of oleandomycin, we collected biological information of abnormalities, associated pathways, cellular/molecular locations, biological functions, related genes/proteins, lipids and common seen animal/experimental models with organized paragraphs from literatures.
What diseases are associated with oleandomycin?
There are no associated biomedical information in the current reference collection.
Possible diseases from mapped MeSH terms on references
We collected disease MeSH terms mapped to the references associated with oleandomycin
PubChem Associated disorders and diseases
What pathways are associated with oleandomycin
There are no associated biomedical information in the current reference collection.
PubChem Biomolecular Interactions and Pathways
Link to PubChem Biomolecular Interactions and PathwaysWhat cellular locations are associated with oleandomycin?
Visualization in cellular structure
Associated locations are in red color. Not associated locations are in black.
Related references are published most in these journals:
| Location | Cross reference | Weighted score | Related literatures |
|---|
What functions are associated with oleandomycin?
Related references are published most in these journals:
| Function | Cross reference | Weighted score | Related literatures |
|---|
What lipids are associated with oleandomycin?
There are no associated biomedical information in the current reference collection.
What genes are associated with oleandomycin?
Related references are published most in these journals:
| Gene | Cross reference | Weighted score | Related literatures |
|---|
What common seen animal models are associated with oleandomycin?
There are no associated biomedical information in the current reference collection.
NCBI Entrez Crosslinks
All references with oleandomycin
Download all related citations| Authors | Title | Published | Journal | PubMed Link |
|---|---|---|---|---|
| Lazarevski G et al. | New oleandomycin 9-oximes. Synthesis, characterization and biological activity. | 1994 | J. Antibiot. | pmid:8175488 |
| Danielson ND et al. | Simple methods for the qualitative identification and quantitative determination of macrolide antibiotics. | 1993 | J Pharm Biomed Anal | pmid:8504183 |
| Hernández C et al. | Characterization of a Streptomyces antibioticus gene cluster encoding a glycosyltransferase involved in oleandomycin inactivation. | 1993 | Gene | pmid:8244027 |
| RodrÃguez AM et al. | Streptomyces antibioticus contains at least three oleandomycin-resistance determinants, one of which shows similarity with proteins of the ABC-transporter superfamily. | 1993 | Mol. Microbiol. | pmid:8326867 |
| Vilches C et al. | Role of glycosylation and deglycosylation in biosynthesis of and resistance to oleandomycin in the producer organism, Streptomyces antibioticus. | 1992 | J. Bacteriol. | pmid:1530845 |
| Kono M et al. | Purification and characterization of macrolide 2'-phosphotransferase type II from a strain of Escherichia coli highly resistant to macrolide antibiotics. | 1992 | FEMS Microbiol. Lett. | pmid:1330822 |
| Il'ina TV | [Assessment of the sensitivity of Neisseria meningitidis strains to benzylpenicillin using the disc diffusion method]. | 1992 | Antibiot. Khimioter. | pmid:1296532 |
| Shevchenko AA et al. | [The interrelation between the formation of oleandomycin and the resistance to it in different strains of Streptomyces antibioticus]. | 1991 Sep-Oct | Mikrobiol. Zh. | pmid:1791777 |
| Terasawa T et al. | In vitro activity of YM133, a new semisynthesized macrolide. | 1991 | Antimicrob. Agents Chemother. | pmid:1929295 |
| Ukhabotina LS and Danilenko VN | [Study of the structure of amplifying sequence of Streptomyces antibioticus]. | 1990 | Antibiot. Khimioter. | pmid:2169231 |